Running part of a workflow multiple times
To run a part of the workflow multiple times, once for each batch unit, add an Iterate control flow element to the workflow. All elements downstream of an Iterate element are run on each batch unit individually, until a Collect and Distribute element is encountered. The parts of the workflow downstream of the Collect and Distribute element are run on all the data together, or in subsets of the data, as configured in the Collect and Distribute element (see Iterate and Collect and Distribute elements).
For example, the workflow in figure 14.80 would run an RNA-Seq Analysis for each sample separately, and then create a single combined report for the set of samples.
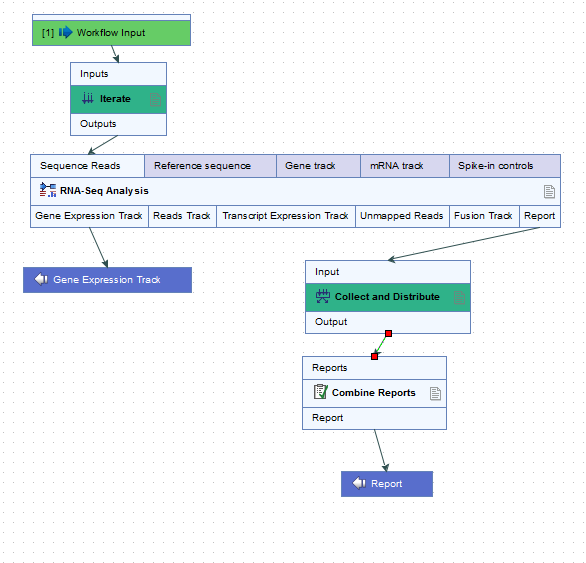
Figure 14.80: The RNA-Seq analysis tool is run once per sample and a single combined report is then generated for the full set of samples.
When running on a CLC Server the iterating parts of the workflow is run as separate jobs. For parallel execution of these iterations, job nodes or grid nodes must be available.
When using metadata table to specify the batch units, you are prompted to specify the column that defines how the samples should be grouped for execution. In figure 14.81, grouping by the column "ID" results in the RNA-Seq Analysis tool being run 8 times, once for each sample. Selecting the "Gender" column instead results in the RNA-Seq Analysis tool being run 2 times, once for each value in that column, male and female. The Combine Reports tool runs once, using information from all samples.
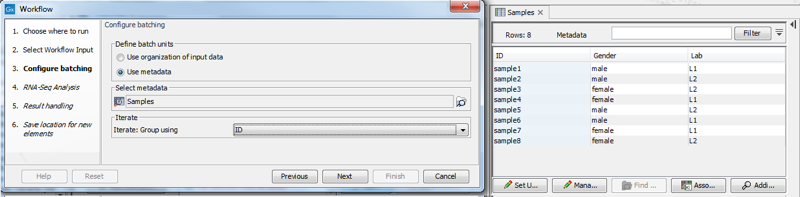
Figure 14.81: With the current selection in the wizard, the RNA-Seq Analysis tool will run 8 times, once for each sample. The Combine Reports tool will run once.
The name of workflow elements can be changed. This changes the text displayed in the wizard when the workflow is run. This can be useful if a workflow contains multiple identical elements (figure 14.82).
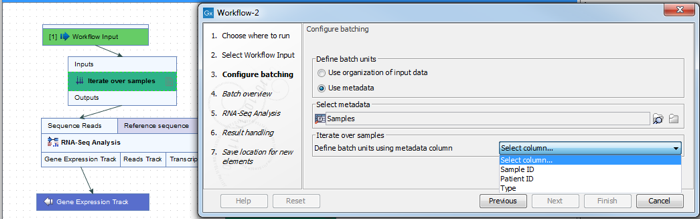
Figure 14.82: The Iterate element can be renamed to change the text that is displayed in the wizard when running the workflow.
