Variant Detection - the outputs
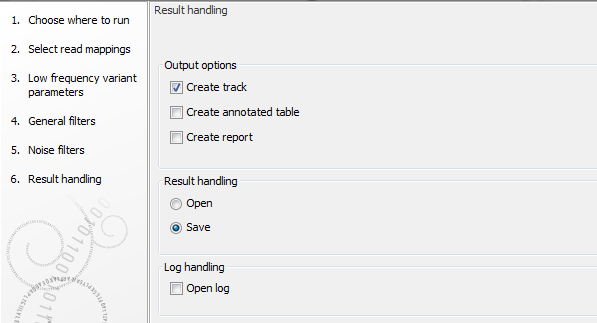
The Variant Detection Tools have the following outputs: a variant track, an annotated variant table and a report (figure 31.15). The report contains information on the estimated error model and, as only the Fixed ploidy and the Low Frequency Variant Detection tool uses an error model, the report is only available for those, and not for the Basic Variant Detection tool.
The outputs are described below. Note that this description can also be applied to variant files that were imported as GVF or VCF files - as described in Import tracks), or downloaded from external databases (e.g. dbSNP, HapMap, or 1000genomes) - as described in Import tracks)).
Subsections