Proteolytic cleavage detection
Given a protein sequence, CLC Genomics Workbench detects proteolytic cleavage sites in accordance with detection parameters and shows the detected sites as annotations on the sequence as well as in a table below the sequence view.
Detection of proteolytic cleavage sites is initiated by:
Tools | Classical Sequence Analysis (![]() ) | Protein Analysis (
) | Protein Analysis (![]() )|
Proteolytic Cleavage (
)|
Proteolytic Cleavage (![]() )
)
If you are connected to a server, the first wizard step will ask you where you would like to run the analysis. Next, you must provide one or more sequences figure 20.25.
Figure 20.25: Choosing a protein sequence for proteolytic cleavage.
In the second dialog, you can select proteolytic cleavage enzymes. Presently, the list contains the enzymes shown in figure 20.26. The full list of enzymes and their cleavage patterns can be seen in Appendix.
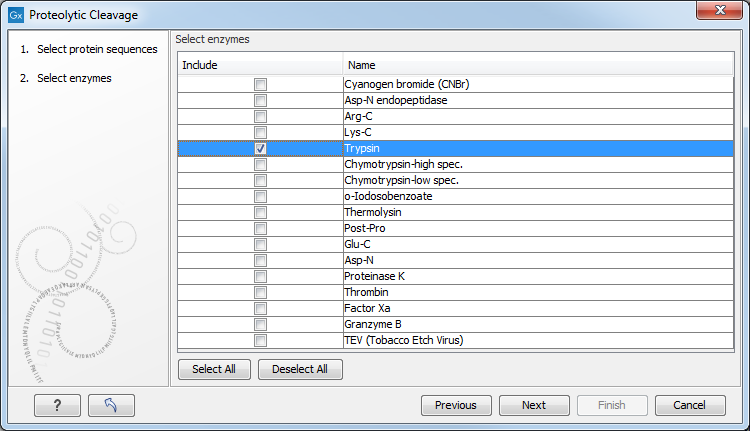
Figure 20.26: Setting parameters for proteolytic cleavage detection.
You can then set parameters for the detection. This limits the number of detected cleavages (figure 20.27).
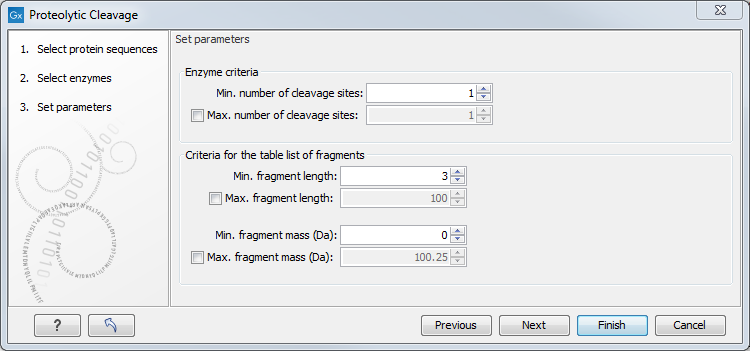
Figure 20.27: Setting parameters for proteolytic cleavage detection.
- Min. and max. number of cleavage sites. Certain proteolytic enzymes cleave at many positions in the amino acid sequence. For instance proteinase K cleaves at nine different amino acids, regardless of the surrounding residues. Thus, it can be very useful to limit the number of actual cleavage sites before running the analysis.
- Min. and max. fragment length Likewise, it is possible to limit the output to only display sequence fragments between a chosen length. Both a lower and upper limit can be chosen.
- Min. and max. fragment mass
The molecular weight is not necessarily directly correlated to the
fragment length as amino acids have different molecular masses. For
that reason it is also possible to limit the search for proteolytic
cleavage sites to mass-range.
For example, if you have one protein sequence but you only want to show which enzymes cut between two and four times. Then you should select "The enzymes has more cleavage sites than 2" and select "The enzyme has less cleavage sites than 4". In the next step you should simply select all enzymes. This will result in a view where only enzymes which cut 2,3 or 4 times are presented.
Click on Finish to launch the analysis. The result of the detection is displayed in figure 20.28.
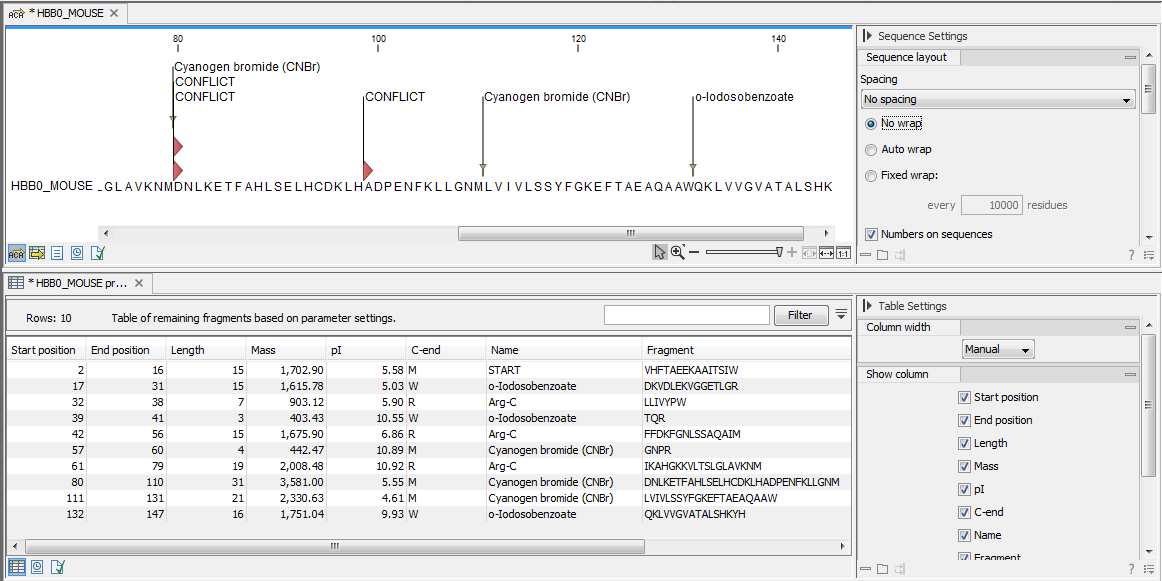
Figure 20.28: The result of the proteolytic cleavage detection.
Depending on the settings in the program, the output of the proteolytic cleavage site detection will display two views on the screen. The top view shows the actual protein sequence with the predicted cleavage sites indicated by small arrows. If no labels are found on the arrows they can be enabled by setting the labels in the "annotation layout" in the preference panel. The bottom view shows a text output of the detection, listing the individual fragments and information on these.
Subsections
