Filter on Custom Criteria
Filter on Custom Criteria takes as input an element, for which data can be displayed as rows in a table view (![]() ), and a set of filter criteria. As many criteria as desired can be defined. The tool outputs an element containing only the rows that satisfy the specified criteria.
), and a set of filter criteria. As many criteria as desired can be defined. The tool outputs an element containing only the rows that satisfy the specified criteria.
Many element types are accepted as input, including:
- Annotation track (
 )
)
- Expression track (
 )
)
- IsomiR table (
 )
)
- miRNA seeds table (
 )
)
- Sequence list (
 )
)
- Statistical comparison table (
 )
)
- Statistical comparison track (
 )
)
- Variant track (
 )
)
To run Filter on Custom Criteria, go to:
Tools | Utility Tools (![]() ) | Filter on Custom Criteria (
) | Filter on Custom Criteria (![]() )
)
After selecting the input element, the criteria to filter on are defined. A criterion is composed of an attribute, a comparison operator, and a value to compare to (figure 37.6).
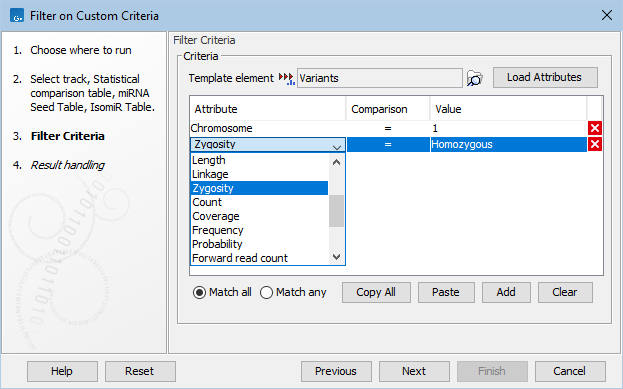
Figure 37.6: Filter criteria to extract the homozygous variants found on chromosome 1. The drop down menu shows some of the attributes populated using the Load Attributes button.
To make defining criteria easier, the Load Attributes button can be used to populate a drop down list of the attributes found in the element selected in the Template element field. By default, the tool's input is preselected. Use the browse (![]() ) button (figure 37.6) to select a different element. We recommend selecting as template a small element containing all necessary attributes, as loading attributes from large elements can take a long time.
) button (figure 37.6) to select a different element. We recommend selecting as template a small element containing all necessary attributes, as loading attributes from large elements can take a long time.
If a template element is not used for populating a drop down list, attributes can be entered by typing directly in the Attribute field.
The Add button adds additional criteria. The X (![]() ) button removes that criterion. The Clear button removes all defined criteria.
) button removes that criterion. The Clear button removes all defined criteria.
Below the criteria definitions, there is an option to choose whether all criteria should be fulfilled (Match all) or if just one needs to be fulfilled (Match any), for a row to be included in the output.
Criteria using attributes not present in the input element are ignored.
Reusing filter criteria
Previously defined filter criteria can be reused in subsequent runs, saving time and making it easy to apply a set of criteria consistently.
Copied filter criteria can be pasted into the 'Filter Criteria' wizard step using the Paste button. Pasting appends the copied criteria to any already defined criteria.
Filter criteria can be copied in two ways:
- The criteria defined in the the 'Filter Criteria' wizard step can be copied using the Copy All button. This can be pasted into a text file for later use.
- The criteria used in previous runs of Filter on Customer Criteria can be copied from the History view (
 ) of the output (figure 37.7).
) of the output (figure 37.7).
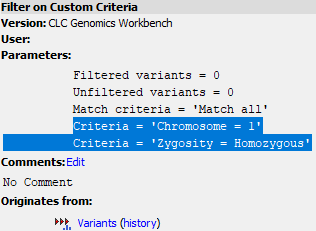
Figure 37.7: The history of an element output by Filter on Custom Criteria includes the criteria used. This can be copied and then pasted into the tool in a subsequent run.
Additional criteria can then be added, and unwanted criteria removed, as described above.
