Extract Reads
Extract Reads takes a stand-alone read mapping or reads track as input and extracts the mapped reads to a reads track or sequence list. All reads or a subset of reads can be extracted. Subsets can be specified based on location relative to a set of regions and/or based on specified characteristics of the reads.
To launch Extract Reads, go to:
Tools | Utility Tools (![]() ) | Extraction (
) | Extraction (![]() ) | Extract Reads (
) | Extract Reads (![]() )
)
Extracting reads based on their location relative to a set of regions can be configured in the 'Select overlap (optional)' wizard step (figure 37.3):
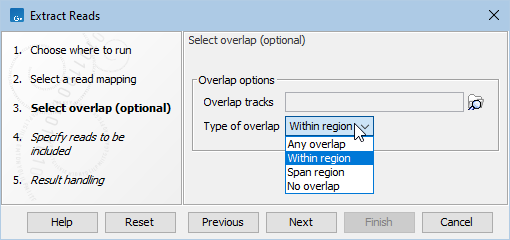
Figure 37.3: Overlap tracks can be used for extracting reads mapped to particular areas of the reference genome.
- Overlap options
- Overlap tracks Extract only reads that overlap one or more regions in the provided overlap track(s). The reference genome of the input and overlap tracks must be compatible.
- Type of overlap (figure 37.4).
- Any overlap. Extract reads that overlap any region.
- Within region. Extract reads fully within an overlap region. Reads overlapping region boundaries are not extracted.
- Span region. Extract reads with residues that align on both sides of an overlap region. For paired reads, fragments that span a region are extracted. The option Only include matching read(s) of read pairs (see below) can be used to solely extract individual reads of a pair that span a region.
- No overlap. Extract reads that do not overlap any region in the provided overlap track(s).
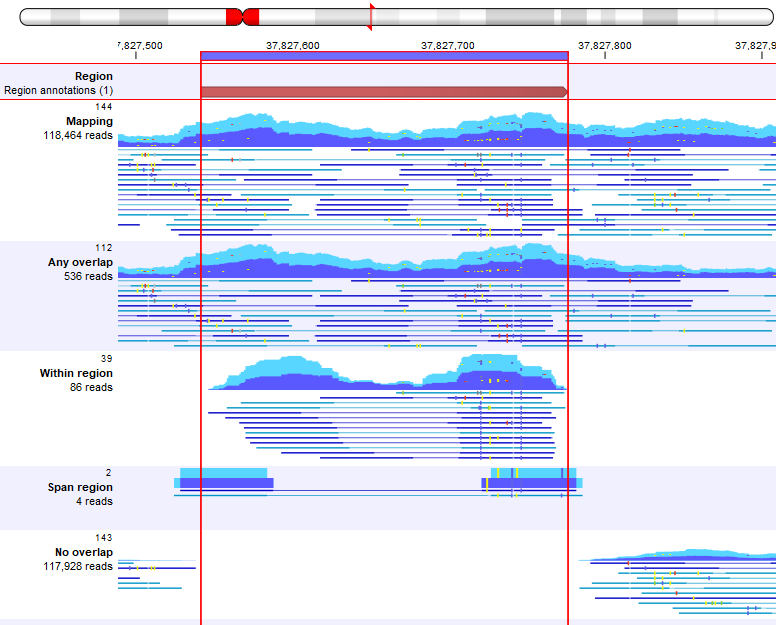
Figure 37.4: A track list including, from top to bottom: an overlap track, a read mapping, and read mappings with extract reads for 'Type of overlap': 'Any overlap', 'Within region', 'Span region', and 'No overlap'.
The nature of the extracted reads can be specified in the 'Specify reads to be included' wizard step (figure 37.5). Note that reads in read mappings are colored according to their characteristics, see Coloring of mapped reads.
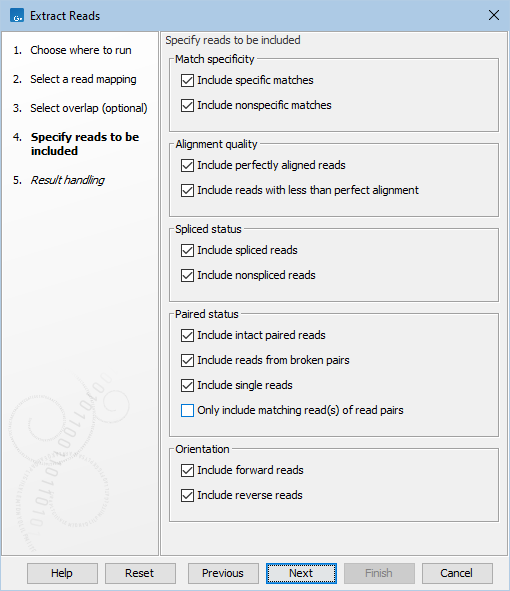
Figure 37.5: Options to include or exclude specific types of reads.
- Match specificity
- Include specific matches Reads that mapped best to just a single position of the reference genome.
- Include non-specific matches Reads that have multiple, equally good alignments to the reference genome. These reads are colored yellow by default in read mappings.
- Alignment quality
- Include perfectly aligned reads Reads where the full read is perfectly aligned to the reference genome. Reads that extend beyond the end of the reference are not considered perfectly aligned, because part of the read does not match the reference.
- Include reads with less than perfect alignment Reads with mismatches, insertions or deletions, or with unaligned ends.
- Spliced status
- Include spliced reads Reads mapped across an intron.
- Include non spliced reads Reads not mapped across an intron.
- Paired status
- Include intact paired reads Paired reads mapped within the specified paired distance.
- Include reads from broken pairs Paired reads where only one of the reads mapped, either because only one read in the pair matched the reference, or because the distance or relative orientation of its mate was wrong.
- Include single reads Reads marked as single reads (as opposed to paired reads). Reads from broken pairs are not included. Reads marked as single reads after trimming paired sequence lists are included.
- Only include matching read(s) of read pairs If only one read of a read pair matches the criteria, then only include the matching read as a broken pair. For example if only one of the reads from the pair is inside the overlap region, then this option only includes the read found within the overlap region as a broken read. When both reads are inside the overlap region, the full paired read is included. Note that some tools ignore broken reads by default.
- Orientation
- Include forward reads Reads mapped in the forward direction.
- Include reverse reads Reads mapped in the reverse direction.
Note that excluding forward or reverse reads will generate broken pairs if reads in pairs are mapped in opposite directions, regardless of the Only include matching read(s) of read pairs option.
