Create RRBS-fragment Track
One common strategy in bisulfite sequencing is to digest genomic DNA with restriction enzymes that cut near CpG islands. For example, the enzyme MspI recognizes and cuts the sequence CCGG. After digestion, these fragments are typically size selected within a defined range before being used for bisulfite sequencing.
Create RRBS-fragment Track generates a feature track with all regions of the reference genome cut by selected restriction enzymes. This track can be used with Call Methylation Levels to restrict statistical testing to these regions of interest.
The following options are available (figure 36.29):
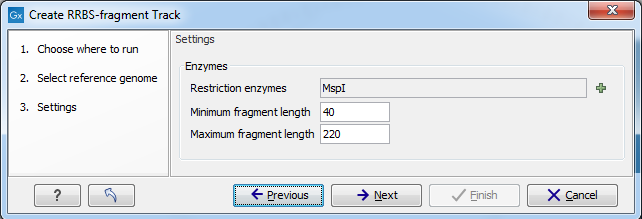
Figure 36.29: Create RRBS-fragment track settings.
- Restriction enzymes The enzymes used in reduced-represention bisulfite sequencing can be selected here, with the most common for the CpG islands, MspI, pre-selected by default.
- Minimum fragment length and Maximum fragment length define the acceptable size range, out of all the possible predicted fragments after full digest of reference DNA, that will be included in the output of the tool.
