Extract sequences
Extract Sequences extracts all the sequences from any of the element types below into a sequence list or into individual sequence elements:
- Alignments (
 )
)
- BLAST result (
 ) For BLAST results, the sequence hits are extracted but not the original query sequence or the consensus sequence.
) For BLAST results, the sequence hits are extracted but not the original query sequence or the consensus sequence.
- BLAST overview tables (
 )
)
- Contigs and read mappings (
 ) For mappings, only the read sequences are extracted. Reference and consensus sequences are not extracted using this tool.
) For mappings, only the read sequences are extracted. Reference and consensus sequences are not extracted using this tool.
- Read mapping tables (
 )
)
- Read mapping tracks (
 )
)
- RNA-Seq mapping results (
 )
)
- Sequence lists (
 ) See further notes below about running this tool on sequence lists.
) See further notes below about running this tool on sequence lists.
If only a subset of the sequences are of interest, create an element containing just this subset first, and then run Extract Sequences on this. See the documentation for the relevant element types for further details. For example, for extracting a subset of a mapping, see Extract parts of a mapping.
Paired reads are extracted in accordance with the read group settings, which are specified during the original import of the reads. If the orientation has since been changed (for example using the Element Info tab for the sequence list), the read group information will be modified and reads will be extracted as specified by the modified read group. The default read group orientation is forward-reverse.
Extracting sequences from sequence lists: As all sequences will be extracted, the main reason to run this tool on a sequence list would be if you wished to create individual sequence elements from each sequence in the list. This is somewhat uncommon. If your aim is to create a list containing a subset of the sequences from another list, this can be done directly from the table view of sequence lists (see Table view of sequence lists), or using Split Sequence List (see Split Sequence List).
Running Extract Sequences
Launch Extract Sequences by going to:
Tools | Classical Sequence Analysis (![]() ) | General Sequence Analysis (
) | General Sequence Analysis (![]() )| Extract Sequences (
)| Extract Sequences (![]() )
)
After selecting the elements to extract sequences from, you are offered the choice of extracting them to individual sequence elements or to a sequence list (figure 18.3). For most data types, a sequence list will be the best choice.
Below these options, the number of sequences that will be extracted is reported.
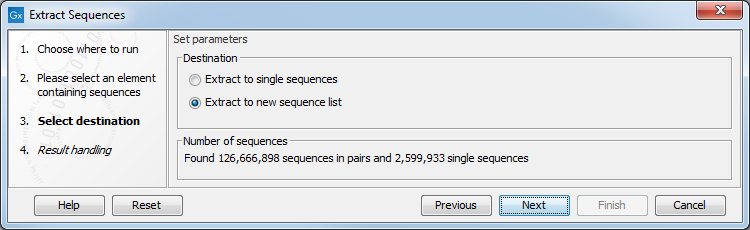
Figure 18.3: Extracted sequences can be put into a new sequence list or split into individual sequence elements.
