MGI/BGI
The MGI/BGI importer is designed to import fastq (.fastq/.fq) files generated by MGI/BGI sequencing technology. Uncompressed files as well as files compressed using gzip (.gz), zip (.zip) or bzip2 (.bz2) can be provided as input. Quality scores are expected to be in the NCBI/Sanger format, see Quality scores in the Illumina platform. The importer processes UMI information from the fastq read headers, see General notes on UMIs.
To launch the MGI/BGI importer, go to:
Import (![]() ) | Other NGS Reads (
) | Other NGS Reads (![]() ) | MGI/BGI (
) | MGI/BGI (![]() ).
).
This opens a dialog where files can be selected and import options specified (figure 7.15).
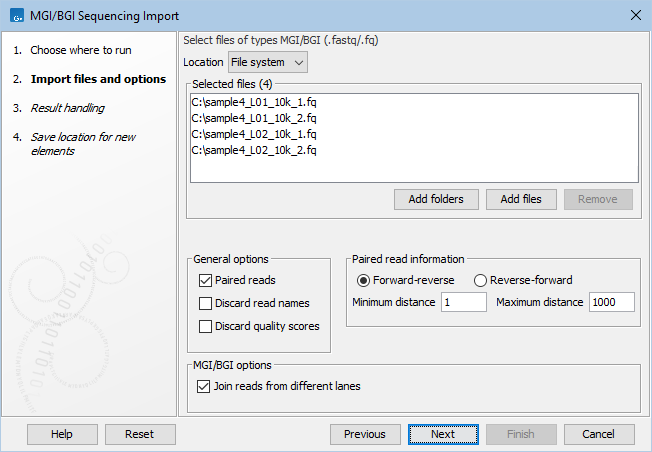
Figure 7.15: Importing data from MGI/BGI.
The General options are:
- Paired reads. Files will be paired up based on their names, which are assumed to contain _1 and _2 (alternatively, _R1 and _R2), respectively. Other than the 1/2 (or the R1/R2), the file names in a pair are expected to be identical. If such a file name format is not used, files will be paired up based on the names of their first read, using one of the following formats:
- The read names end with /1 and /2, for example
@sample1/1and@sample1/2. - The read names contain a space followed by 1 or 2, for example
@sample1 1:NNNand@sample1 2:NNN.
Under Paired read information:
- Choose the orientation of the paired reads, either Forward-reverse or Reverse-forward.
- Specify the insert sizes by setting Minimum distance and Maximum distance. Data sets with different insert sizes should be imported separately, with the correct minimum and maximum distance.
Read more about handling paired data in General notes on handling paired data.
- The read names end with /1 and /2, for example
- Discard read names. Read names can be discarded to save disk space without affecting analysis results. Keeping read names can be useful in some circumstances, such as when inspecting sequence list contents or when working downstream with subsets of sequences.
- Discard quality scores. Quality scores are visible in read mappings and are used by various tools, e.g. for variant detection. If quality scores are not relevant, use this option to discard them and reduce disk space and memory consumption.
The MGI/BGI options are:
- Join reads from different lanes.
When checked, fastq files from the same sequencing run but from different lanes are imported as a single sequence list.
Lane information is expected in the filenames as "_L<digits>", e.g. "L001" for lane 1. If this patterns occurs more than once in a filename, the last occurrence in the name is used. For example, for a filename "myFile_L001_L1.fastq" the lane information is assumed to be L1.
