Tracks
Tracks provide a unified framework for the visualization, comparison and analysis of genome-scale studies such as whole-genome sequencing or exome resequencing projects. Information in tracks is tied to a genomic position. Each track type supports a particular type of data, with functionality relevant to the type of data available. Details about the different types of tracks are provided in Track types.
When the mouse cursor is hovered over items in the graphical view of a track, information about that item and about the location the cursor is hovered over is displayed in several places (figure 27.1):
- Just below the coordinates near the top. Where relevant, the following are also displayed:
- The item name.
- The strand the item is located on.
- For items comprised of multiple parts, e.g. mRNAs containing multiple exons, the number of the part that the cursor is hovered over.
- The position of the cursor relative to the start of the item.
- In the View Area: A tooltip containing information about the item.
- In the lower right corner of the Workbench frame, as relevant for the item type, including:
- The item type.
- The item name.
- The position of the cursor relative to the start of the item.
- For items comprised of multiple parts, e.g. mRNAs containing multiple exons, the number of the part that the cursor is hovered over.
- The strand the item is located on.
- The position of the cursor relative to the reference.
- Other information relevant to the item. For example, for variants, the variant type and its size.
Compatible tracks can be stacked in a Track List, supporting comparative analysis (figure 27.1). See Track lists for details about creating and working with Track Lists.
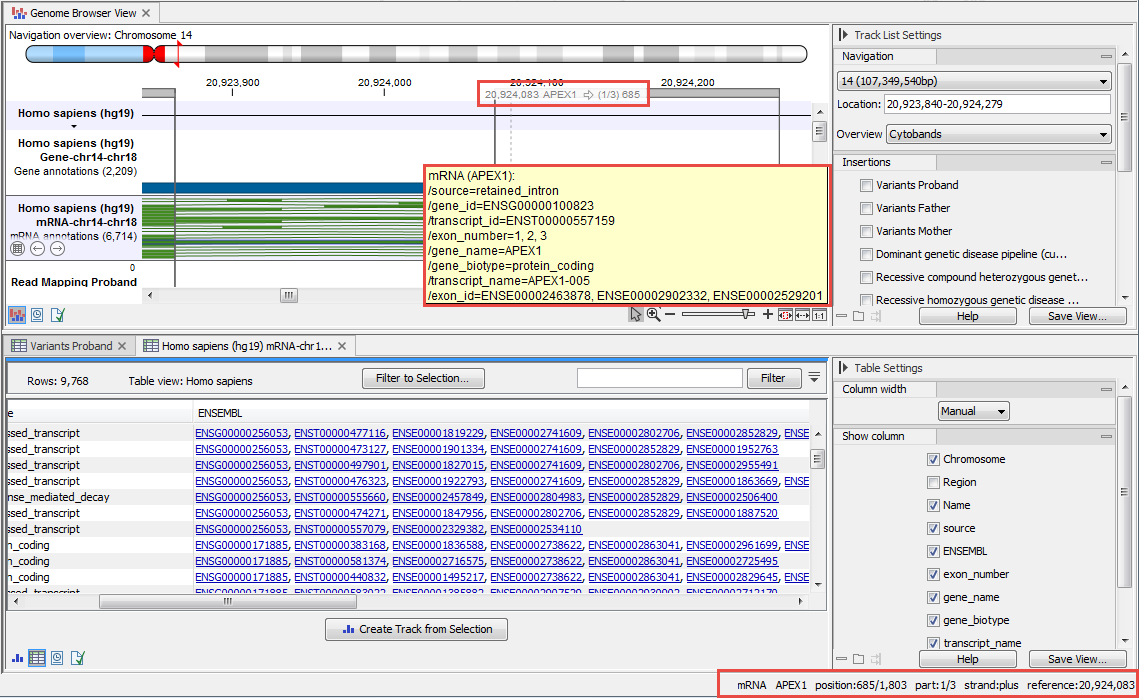
Figure 27.1: Compatible tracks have been added to a Track List and the cursor is hovered over an item in one of the tracks, revealing information about that item in three places: just under the coordinates near the top, in a tooltip near the item itself, and in the lower right corner of the Workbench frame.
Subsections
- Track types
- Track lists
- Working with tracks
- Reference data as tracks
- Convert tracks
- Graph tracks
- Merge tracks
- Modify tracks
