Variant Detection tools
CLC Genomics Workbench offers three tools for detecting variants.
- Fixed Ploidy Variant Detection (
 ) - described in detail in section Fixed Ploidy Variant Detection
) - described in detail in section Fixed Ploidy Variant Detection
- Low Frequency Variant Detection (
 ) - described in detail in section Low Frequency Variant Detection
) - described in detail in section Low Frequency Variant Detection
- Basic Variant Detection (
 ) - described in detail in section Basic Variant Detection
) - described in detail in section Basic Variant Detection
They are designed for the analysis of different types of samples and they differ in their underlying assumptions about the data, and hence in their assessments of when there is enough information in the data for a variant to be called. An overview of these differences is given in figure 31.1.
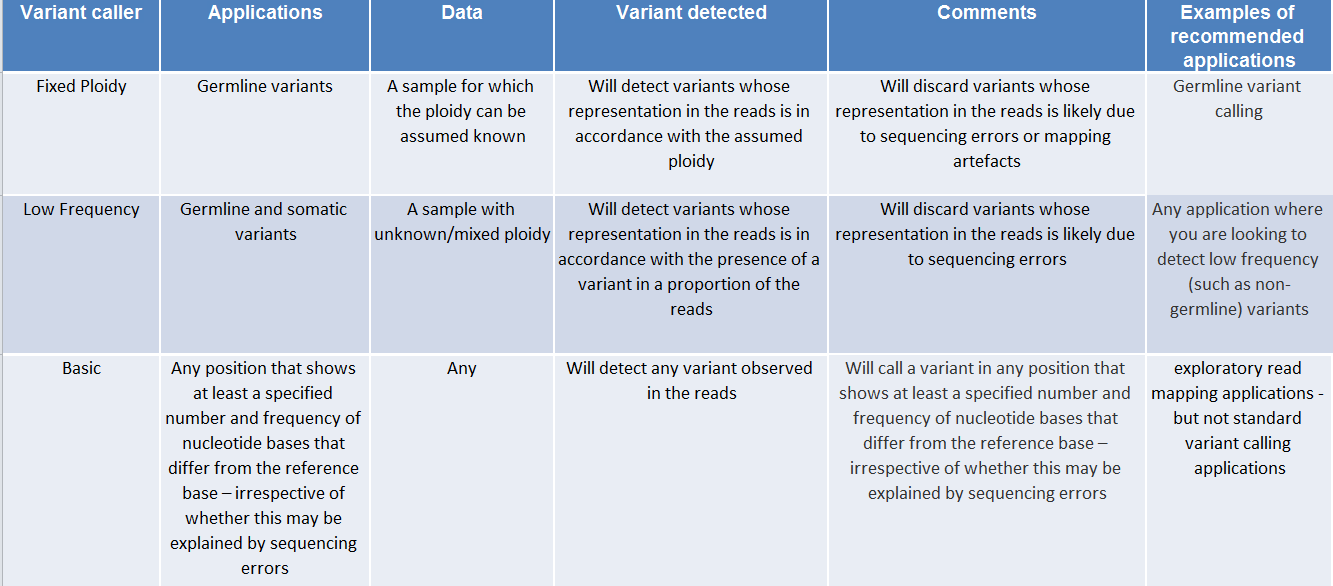
Figure 31.1: An overview of the variant detection tools.
To run a variant detection tool, go to:
Tools | Resequencing Analysis (![]() ) | Variant Detection (
) | Variant Detection (![]() )
)
and choose the appropriate tool. In the first dialog of each detection tool, you are asked to specify the reads track or read mapping to analyze. The user is next asked to set the parameters that are specific for the variant detection tool. The three tools, their assumptions, and the tool-specific parameters are described later in their respective sections.
All variant detection tools will call:
- SNVs - single nucleotide variants
- MNVs - neighboring SNVs, where there is evidence they occur together
- small to medium-sized insertions and deletions - insertions and deletions fully represented within a single read
- replacements - neighboring SNVs and insertions or deletions
Subsections
- Differences in the variants called by the different tools
- How the variant detection tools work
- Detailed information about overlapping paired reads
