Update Sequence Attributes in Lists
Update Sequence Attributes in Lists updates information about sequences in a Sequence List. Information can be added to existing attributes and new attribute types can be added.
The tool takes attribute information from an Excel file (.xls/xlsx), a comma separated text file (.csv), or a tab separated text file (.tsv). Each sequence is updated with the relevant information by matching the content of a particular column in the file, specified when launching the tool, with the contents of a column of the same name in the Sequence List.
The columns to take information from are specified when launching the tool. Column names are used as attribute names. If a column name matches an existing attribute in the Sequence List, the information from that column can be added to the existing attribute (details below). When a column name does not match an existing attribute, a new attribute is added to the Sequence List.
Additional notes:
- This tool is recommended when updating information for many sequences. However, attributes can also be updated individually, either directly in the Table view of the Sequence List (see Table view of sequence lists), or, by opening the sequence from the Sequence List and editing attributes in its Element info view. Opening a sequence can be done from the Sequence List view (right-click on the sequence name and choose "Open Sequence") or from the Table view (right-click in the row for that sequence and choose the option "Open This Sequence").
- Attributes relating to characteristics of the sequence itself, such as its length or the start of the sequence, cannot be updated using this tool, nor by directly editing the Sequence List.
To launch the Update Sequence Attributes in Lists tool, go to:
Tools | Utility Tools (![]() ) | Sequence Lists (
) | Sequence Lists (![]() ) |Update Sequence Attributes in Lists (
) |Update Sequence Attributes in Lists (![]() )
)
and select one or more Sequence Lists of the same type (nucleotide or peptide) as input (figure 37.32).
Note: Sequences in all inputs provided will be worked upon as a single entity. A single Sequence List containing all sequences is output.
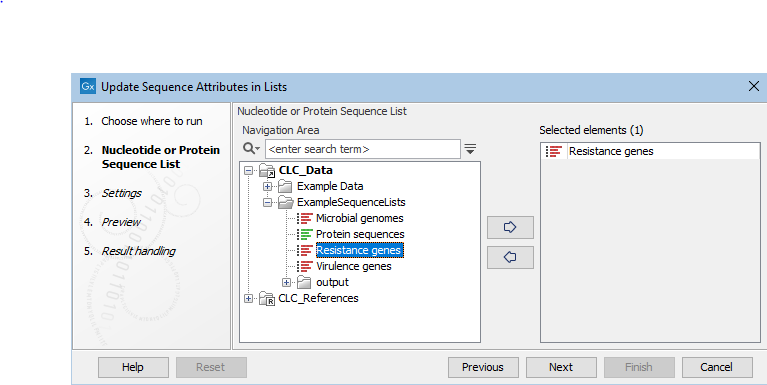
Figure 37.32: Select one or more Sequence Lists as input.
In the Settings wizard step, the file containing attribute information is specified, along with details about how to handle that information (figure 37.33).
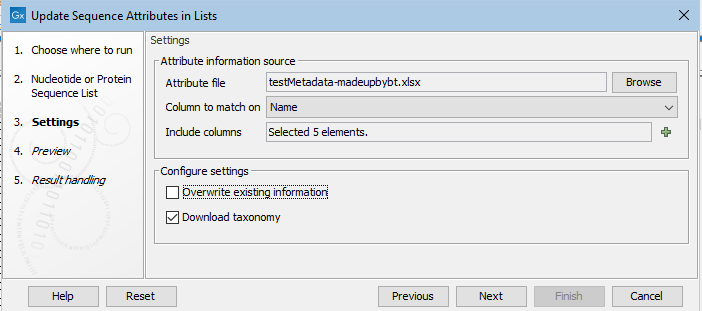
Figure 37.33: Information in the attribute file will be matched with the relevant sequence based on contents of the Name column in the file and in the Sequence List. Five columns containing relevant attribute information have been selected. The option to overwrite existing information has been left unchecked.
Attribute information source fields
- Attribute file Select an Excel file (
.xls/xlsx), a comma separated text file (.csv), or a tab separated text file (.tsv) containing attribute information. Column names are used as attribute names, so a header row is required. One column in the file must contain information that can be matched with information already present in the Sequence List (see "Column to match on", below). - Column to match on Specify the column in the attribute file to use to match each row with the relevant sequence(s) in the Sequence List. When a value in this column matches a value in the column of the same name in the Sequence List, information from that row in the file is added to the attribute information for that sequence. Only information from specified columns will be added (see "Include columns", below.)
When matching based on sequence names, the column in the file containing the names must be called
Name. - Include columns Select the columns in the file containing the information to be updated or added to the Sequence List as well as the column specified in the "Column to match on" field.
When the name of a column does not match existing attribute name in the Sequence List, a new attribute will be added.
Configure settings checkboxes
- Overwrite existing information When this option is checked, existing sequence attribute values will be overwritten by values for the corresponding attributes in the attribute file. When no corresponding value is present in the attributes file, no change is made to the value in the Sequence List.
When left unchecked, existing attribute values in the Sequence List are not overwritten with new information from the file.
- Download taxonomy Check this box to download a 7-step taxonomy from the NCBI into an attribute called "Taxonomy". To use this option, there must be a column in the attributes file called
TaxIDcontaining valid taxonomic identifiers. See the "Column headings and value validation" section below for further details.The "Taxonomy" attribute will be listed in the Preview wizard step, alongside the columns selected for inclusion.
The result of the choices made in the Settings step are reflected in the Preview wizard step (figure 37.34). In the upper pane is a list of the attribute types to be updated or added, as well as the attribute to be used to match sequences with the relevant information. How particular columns will be handled is indicated in the "Content handling" column, including whether validation will be applied. The columns subject to validation checks are described later in this section.
Shown in the lower pane is a small subset of the incoming information from the attribute file, based on the choices made in the Settings wizard step. Click on the "Previous" button to go back to that step if anything needs to be adjusted.
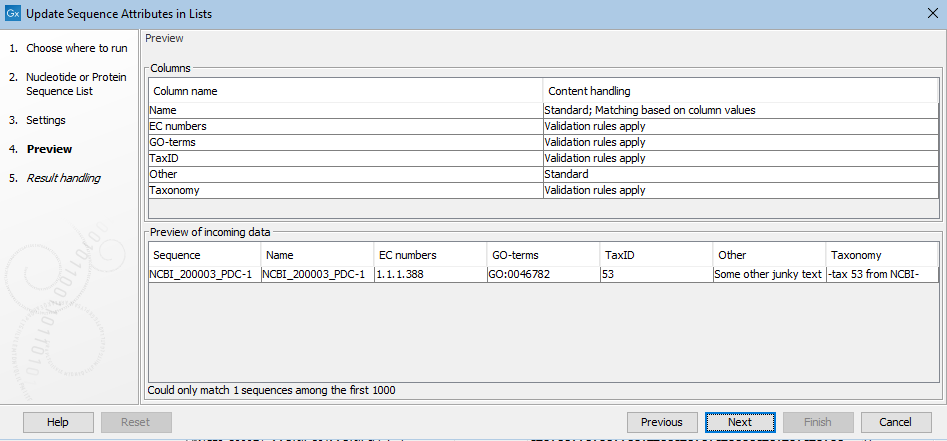
Figure 37.34: The Preview wizard steps shows information about how columns from the attribute file will be handled, and whether any problems were detected. Where validation checks are carried out, if any had failed, a yellow exclamation mark in the bottom pane would be shown for that column. Here, all entries pass. The "Other" column is not subject to validation checks. Only one sequence in the list is being updated in this example.
Column headings and value validation
Certain column names are recognized by the software and validation rules are applied to these. When the contents pass the validation checks, entries in those columns may be further processed.
In most cases, this further processing involves adding hyperlinks to online data resources. However, the contents of columns with the following names trigger different handling:
- TaxID When valid taxonomic identifiers are found in a column called
TaxID, and the Download taxonomy checkbox was checked in the Settings wizard step, then a 7-step taxonomy is downloaded from the NCBI.Examples of valid identifiers for TaxID attribute are those found in
/db_xref="taxonfields in Genbank entries. For example, for/db_xref="taxon:5833, the expected value in the TaxID column would be5833.If a given sequence has an value already set for the Taxonomy attribute, then that existing value remains in place unless the "Overwrite existing information" box was checked in the Settings wizard step.
- Gene ID The following identifiers in a Gene ID column are added as attribute values and hyperlinked to the relevant online database:
- Ensembl Gene IDs
- HUGO Gene IDs
- VFDB Gene IDs
Multiple identifiers in a given cell, separated by commas, will be added as multiple Gene ID attributes for the relevant sequence. If any one of those identifiers is not recognized as one of the above types, then none will be hyperlinked.
Other columns where contents are validated are those with the headings listed below. If a value in such a column cannot be validated, it is not added nor used to update attributes.
If you wish to add information of this type but do not want this level of validation applied, use a heading other than the ones listed below.
- COG-terms COG identifiers
- Compound AROs Antibiotic Resistance Ontology identifiers
- Compound Class AROs Antibiotic Resistance Ontology identifiers
- Confers-resistance-to ARO Antibiotic Resistance Ontology identifiers
- Drug ARO Antibiotic Resistance Ontology identifiers
- Drug Class ARO Antibiotic Resistance Ontology identifiers
- EC numbers EC identifiers
- GenBank accession Genbank accession numbers
- Gene ARO Antibiotic Resistance Ontology identifiers
- GO-terms Gene Ontology (GO) identifiers
- KO-terms KEGG Orthology (KO) identifiers
- Pfam domains PFAM domain identifiers
- Phenotype ARO Antibiotic Resistance Ontology identifiers
- PubMed IDs Pubmed identifiers
- TIGRFAM-terms TIGRFAM identifiers
- Virulence factor ID Virulence Factors of Pathogenic Bacteria identifiers
