Mapping table
When several reference sequences are used or you are performing de novo assembly with the reads mapped back to the contig sequences, all your mapping data will be accessible from a table (An example of a mapping table for a de novo assembly is shown in figure 30.32.
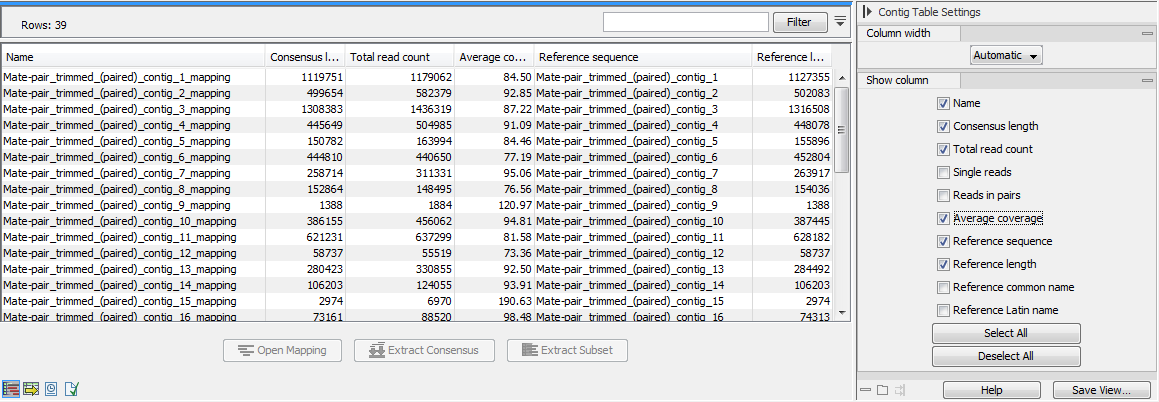
Figure 30.32: The mapping table.
The information included in the table is:
- Name. When mapping reads to a reference, this will be the name of the reference sequence.
- Consensus length. The length of the consensus sequence. Subtracting this from the length of the reference will indicate how much of the reference that has not been covered by reads.
- Total read count. The number of reads. Reads with multiple hits on different reference sequences are placed according to your input for Non-specific matches
- Single reads and Reads in pair. Total number of reads, single and/or in pair.
- Average coverage. This is simply summing up the bases of the aligned part of all the reads divided by the length of the reference sequence.
- Reference sequence. The name of the reference sequence.
- Reference length. The length of the reference sequence.
- Reference common name and
- Reference latin name. Name, common name and Latin name of each reference sequence.
At the bottom of the table there are three buttons that can be used to open or extract sequences. Select the relevant rows before clicking on the buttons:
- Open Mapping. Opens the read mapping for visual inspection. You can also open one mapping simply by double-clicking in the table.
- Extract Consensus/Contigs. For de novo assembly results, the contig sequences will be extracted. For results when mapping against a reference, the Extract Consensus tool will be used (see Extract consensus sequence).
- Extract Subset. Creates a new mapping table with the mappings that you have selected.
You can export the table in Excel format.
