Showing a track in a table
All tracks containing annotations (including variants) can be opened in a table. it is usually helpful to open the table in split view so that both the track and the table can be seen at the same time. In particular, because track and table are connected, it becomes possible to navigate and zoom the track by selecting successively the different rows in the table. Similarly, making a selection in the track view will select the corresponding row in the table. Buttons in the variant table (figure 27.9) view can filter the table to show only the region selected in the track. One can also create a new track from the rows selected in the table.
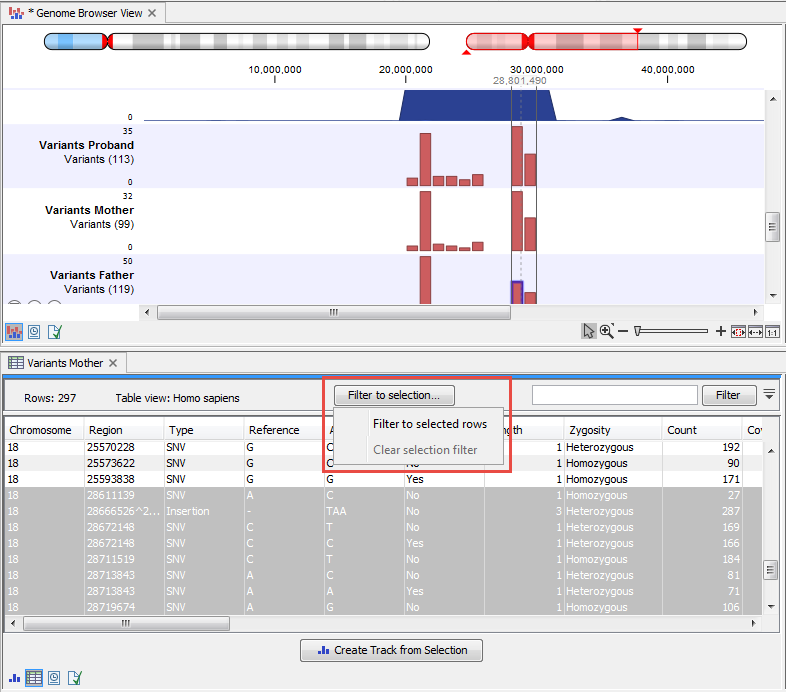
Figure 27.9: Filter the table to display only the rows selected in the track using the menu highlighted in red. It is also possible to create a new track from the selected rows.
To open a track as a table in split view, press Ctrl (![]() on Mac) while you click the table button at the bottom of the track view. You can also right-click on the track tab, and select "Show View | Table".
on Mac) while you click the table button at the bottom of the track view. You can also right-click on the track tab, and select "Show View | Table".
The table will have one row for each annotation, and the columns will reflect its information content. Figure 27.10 shows an example of a variant database track that is presented in a table.
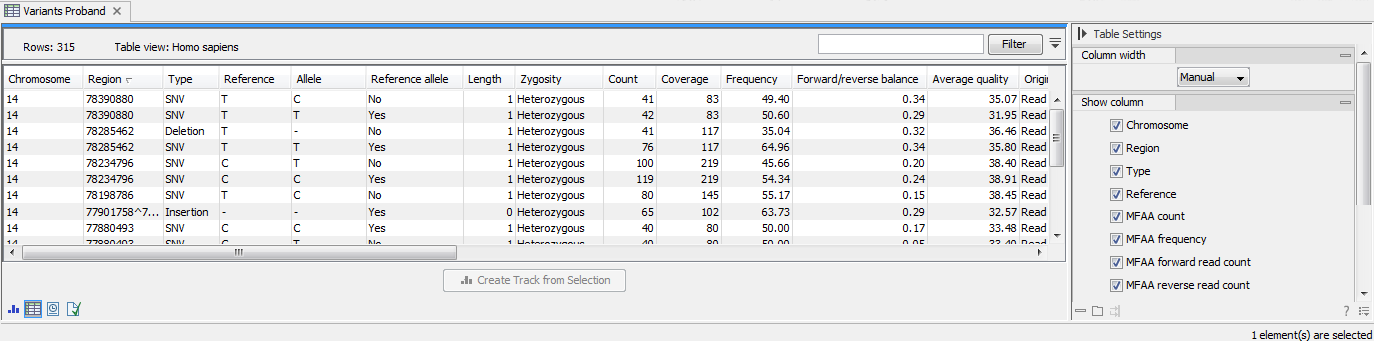
Figure 27.10: Showing a variant track in a table view.
You can use the table to sort, filter and select annotations (see Working with tables). Please note that there are two additional options for filtering on overlaps in the "Region" column
When selecting a row in the table the graphical view will jump to this position on the genome. Please note that table filtering only affects the table. The track itself remains unaffected and keeps all annotations. If you also wish to filter tracks in the graphical view, the Annotate and Filter tools (see Annotate and Filter) can be used instead.
At the bottom of the table a button labeled Create Track from Selection is available. This function can be used to create tracks showing only a subset of the data and annotations. Select the relevant rows in the table and click the button to create a new track that only includes the selected subset of the annotations. This function is particularly useful when used in combination with the filter.
