Import high-throughput sequencing data
CLC Genomics Workbench has dedicated tools for importing data from the following High-throughput sequencing systems:
- QIAGEN GeneReader
- Illumina Genome Analyzer, Nextseq, HiSeq and MiSeq
- PacBio
- Ion Torrent
It is also possible to import mapped data in SAM/BAM format, meaning that alignments of Complete Genomics data can be imported using the SAM/BAM importer.
An importer for Roche 454 sequencing data is available in the Legacy Tools folder. While we do not support the import of files in Complete Genomics master VAR file format, these files can be converted to VCF using the tools provided by Complete Genomics, and later imported in the workbench as VCF.
Having dedicated tools for importing NGS data helps standardize the data so that downstream analyses and visualization work seamlessly with all sequencing platforms. In case a sequence list was not imported with the right tool, it is possible to edit "Read Group" information in the "Element Info" view: choose from the drop-down menu the sequencing platform that was used to generate the data (figure 6.7) and click OK.
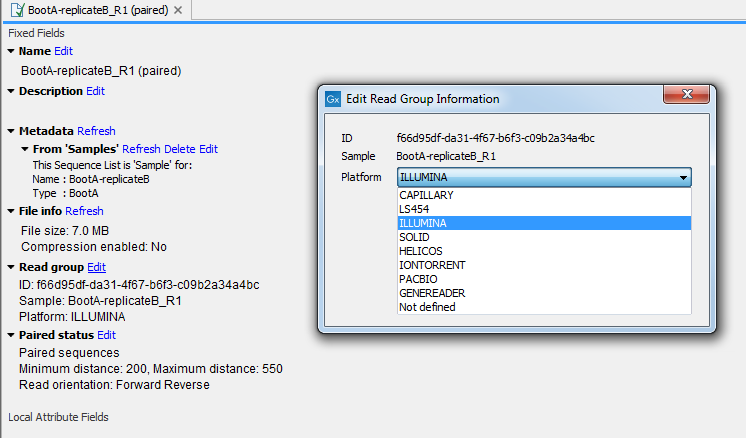
Figure 6.7: Editing the platform that was used to generate the data in the "Element Info" view.
Clicking on the Import (![]() ) button in the top toolbar will bring up a list of the supported data types as shown in figure 6.8.
) button in the top toolbar will bring up a list of the supported data types as shown in figure 6.8.
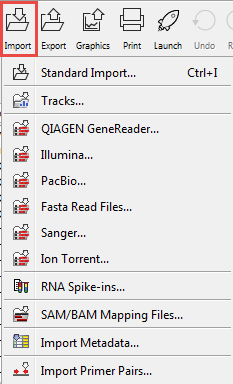
Figure 6.8: Choosing what kind of data you wish to import.
Select the appropriate format and then fill in the information as explained in the following sections.
Subsections
- QIAGEN GeneReader
- Illumina
- PacBio
- Fasta read files
- Sanger sequencing data
- Ion Torrent
- General notes on handling paired data
- SAM and BAM mapping files
