Annotate with Flanking Sequence
In some situations, it is useful to see a variant in the context of the bases of the reference sequence. This information can be added using the Annotate with Flanking Sequence tool:
Toolbox | Resequencing Analysis (![]() ) | Variant Annotation (
) | Variant Annotation (![]() ) | Annotate with Flanking Sequence
) | Annotate with Flanking Sequence
This opens a dialog where you can select a variant track (![]() ) to be annotated.
) to be annotated.
Clicking Next will display the dialog shown in figure 27.32
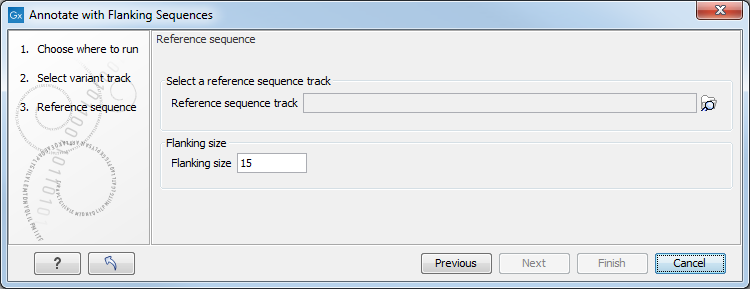
Figure 27.32: Specifying a reference sequence and the amount of flanking bases to include.
Select a sequence track that should be used for adding the flanking sequence, and specify how large the flanking region should be.
The result will be a new track with an additional column for the flanking sequence formatted like this: CGGCT[T]AGTCC with the base in square brackets being the variant allele.
