Signal peptide prediction parameter settings
It is possible to set different options prior to running the analysis (see figure 16.1). An organism type should be selected. The default is eukaryote.
- Eukaryote (default)
- Gram-negative bacteria
- Gram-positive bacteria
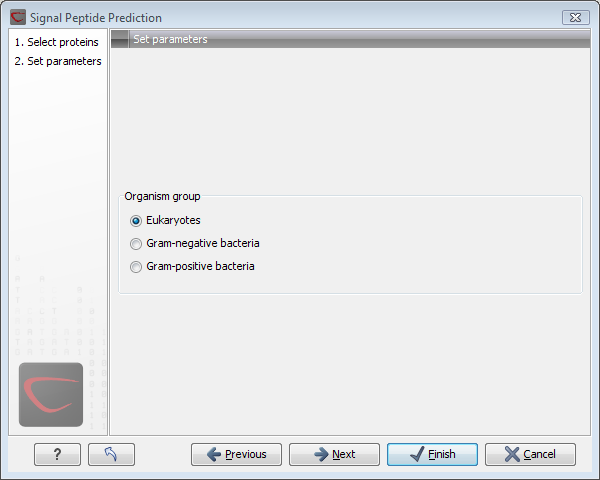
Figure 16.1: Setting the parameters for signal peptide prediction.
You can perform the analysis on several protein sequences at a time. This will add annotations to all the sequences and open a view for each sequence if a signal peptide is found. If no signal peptide is found in the sequence a dialog box will be shown.
The predictions obtained can either be shown as annotations on the sequence, listed in a table or be shown as the detailed and full text output from the SignalP method. This can be used to interpret borderline predictions:
- Add annotations to sequence
- Create table
- Text
