Selecting, sorting and filtering enzymes
Clicking Next lets you define which enzymes to use as basis for finding restriction sites on the sequence. At the top, you can choose to Use existing enzyme list. Clicking this option lets you select an enzyme list which is stored in the Navigation Area. See section Restriction enzyme list for more about creating and modifying enzyme lists.Below there are two panels:
- To the left, you see all the enzymes that are in the list select above. If you have not chosen to use an existing enzyme list, this panel shows all the enzymes available 19.5.
- To the right, there is a list of the enzymes that will be used.
Select enzymes in the left side panel and add them to the right
panel by double-clicking or clicking the Add button
(![]() ). If you e.g. wish to use EcoRV and BamHI, select these
two enzymes and add them to the right side panel.
). If you e.g. wish to use EcoRV and BamHI, select these
two enzymes and add them to the right side panel.
If you wish to use all the enzymes in the list:
Click in the panel to the left | press Ctrl + A
(![]() + A on Mac) | Add (
+ A on Mac) | Add (![]() )
)
The enzymes can be sorted by clicking the column headings, i.e. Name, Overhang, Methylation or Popularity. This is particularly useful if you wish to use enzymes which produce e.g. a 3' overhang. In this case, you can sort the list by clicking the Overhang column heading, and all the enzymes producing 3' overhangs will be listed together for easy selection.
When looking for a specific enzyme, it is easier to use the Filter. If you wish to find e.g. HindIII sites, simply type HindIII into the filter, and the list of enzymes will shrink automatically to only include the HindIII enzyme. This can also be used to only show enzymes producing e.g. a 3' overhang as shown in figure 19.51.
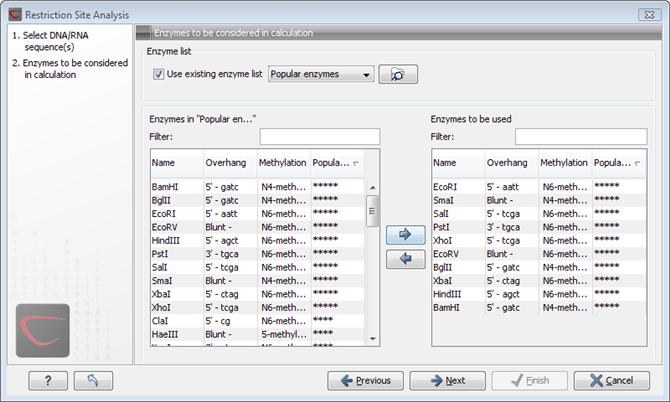
Figure 19.41: Selecting enzymes.
If you need more detailed information and filtering of the enzymes, either place your mouse cursor on an enzyme for one second to display additional information (see figure 19.52), or use the view of enzyme lists (see Restriction enzyme list).
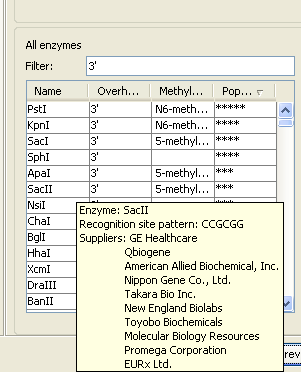
Figure 19.42: Showing additional information about an enzyme like recognition sequence or a list of commercial vendors.
Footnotes
- ... available19.5
- The CLC Genomics Workbench comes with a standard set of enzymes based on http://www.rebase.neb.com. You can customize the enzyme database for your installation, see Restriction enzymes database configuration
