Advanced search
As a supplement to the Quick search you can use the more advanced search:
Edit | Local Search (![]() )
)
or Ctrl + F (![]() + F on Mac)
+ F on Mac)
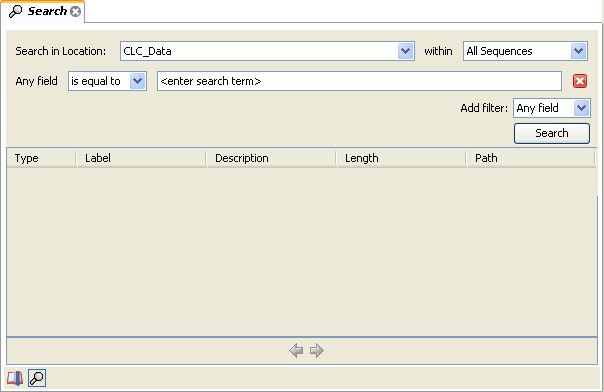
This will open the search view as shown in figure 3.20
The first thing you can choose is which location should be searched. All the active locations are shown in this list. You can also choose to search all locations. Read more about Data structure.
Furthermore, you can specify what kind of elements should be searched:
- All sequences
- Nucleotide sequences
- Protein sequences
- All data
When searching for sequences, you will also get alignments, sequence lists etc as result, if they contain a sequence which match the search criteria.
Below are the search criteria. First, select a relevant search filter in the Add filter: list. For sequences you can search for
- Name
- Length
- Organism
See this list for more information on individual search terms.
For all other data, you can only search for name.
If you use Any field, it will search all of the above plus the following:
- Description
- Keywords
- Common name
- Taxonomy name
To see this information for a sequence, switch to the Element Info (![]() ) view.
) view.
For each search line, you can choose if you want the exact term by selecting "is equal to" or if you only enter the start of the term you wish to find (select "begins with").
An example is shown in figure 3.21.
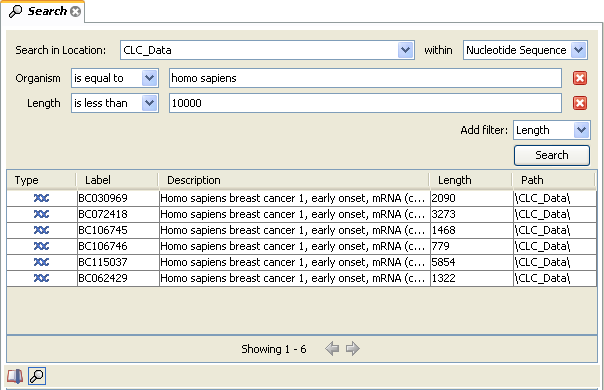
Figure 3.21: Searching for human sequences shorter than 10,000 nucleotides.
This example will find human nucleotide sequences (organism is Homo sapiens), and it will only find sequences shorter than 10,000 nucleotides.
Note that a search can be saved (![]() ) for later use. You do
not save the search results - only the search parameters. This means
that you can easily conduct the same search later on when your data
has changed.
) for later use. You do
not save the search results - only the search parameters. This means
that you can easily conduct the same search later on when your data
has changed.