Secondary structure prediction
An important issue when trying to understand protein function is to know the actual structure of the protein. Many questions that are raised by molecular biologists are directly targeted at protein structure. The alpha-helix forms a coiled rodlike structure whereas a beta-sheet show an extended sheet-like structure. Some proteins are almost devoid of alpha-helices such as chymotrypsin (PDB_ID: 1AB9) whereas others like myoglobin (PDB_ID: 101M) have a very high content of alpha-helices.
With CLC Genomics Workbench one can predict the secondary structure of proteins very fast. Predicted elements are alpha-helix, beta-sheet (same as beta-strand) and other regions.
Based on extracted protein sequences from the protein databank (http://www.rcsb.org/pdb/) a hidden Makov model (HMM) was trained and evaluated for performance. Machine learning methods have shown superior when it comes to prediction of secondary structure of proteins [Rost, 2001]. By far the most common structures are Alpha-helices and beta-sheets which can be predicted, and predicted structures are automatically added to the query as annotation which later can be edited.
In order to predict the secondary structure of proteins:
Select a protein sequence | Toolbox in the Menu
Bar | Classical Sequence Analysis (![]() ) | Protein Analysis (
) | Protein Analysis (![]() )|
Predict secondary structure (
)|
Predict secondary structure (![]() )
)
or right-click a protein sequence | Toolbox |
Classical Sequence Analysis (![]() ) | Protein Analysis (
) | Protein Analysis (![]() )| Predict
secondary structure (
)| Predict
secondary structure (![]() )
)
This opens the dialog displayed in figure 16.18:
Figure 16.18: Choosing one or more protein sequences for secondary structure prediction.
If a sequence was selected before choosing the Toolbox action, this sequence is now listed in the Selected Elements window of the dialog. Use the arrows to add or remove sequences or sequence lists from the selected elements.
You can perform the analysis on several protein sequences at a time. This will add annotations to all the sequences and open a view for each sequence.
Click Next if you wish to adjust how to handle the results. If not, click Finish.
After running the prediction as described above, the protein sequence will show predicted alpha-helices and beta-sheets as annotations on the original sequence (see figure 16.19).
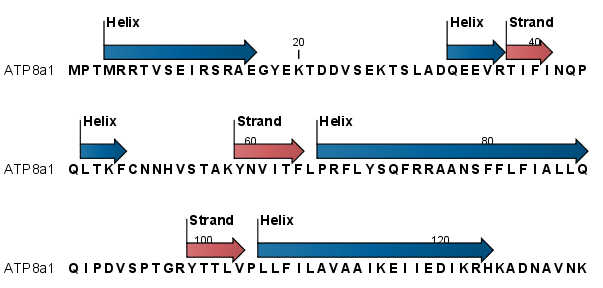
Figure 16.19: Alpha-helices and beta-strands shown as annotations on the sequence.
Each annotation will carry a tooltip note saying that the
corresponding annotation is predicted with CLC Genomics Workbench.
Additional notes can be added through the Edit Annotation
(![]() ) right-click mouse menu. Removing
annotations.
) right-click mouse menu. Removing
annotations.
Undesired alpha-helices or beta-sheets can be removed through the
Delete Annotation (![]() ) right-click mouse
menu. Removing annotations.
) right-click mouse
menu. Removing annotations.
