Minimum Spanning Trees
A minimum spanning tree (MST) is a tree connecting all nodes in a graph, in a way such that the sum of edge lengths is minimized, see figure 10.4.
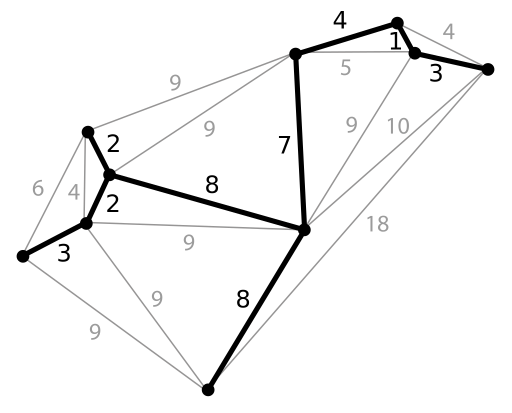
Figure 10.4: Minimum Spanning Tree construction. (Public domain image from Wikimedia: Minimum spanning tree.svg.)
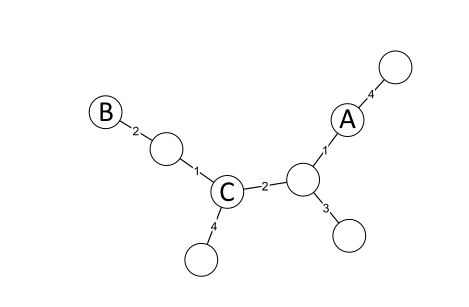
Figure 10.5: Minimum Spanning Tree distances.
Edges in the MST show the strongest local links, not actual cumulative distances. This means you cannot sum the distances of multiple edges to get the true distance between two nodes that are not directly connected - the MST only guarantees minimal total connectivity, not accurate pairwise distances. Consider the tree in figure 10.5. The distance between A and B is not necessarily larger than the distance from A to C. But we do know that the distance between two nodes is greater or equal to the largest edge connecting them (e.g. the distance between A and B is at least two, and the distance between A and C is at least two).
MSTs are often used to visualize relationships between strains or isolates. But note that MSTs are not unique - there are often many possible trees, especially for the classic 7-gene schemes, where there are only a very limited number of possible edge lengths. In order to break the ties when constructing the tree, our MST implementation favors creating connections to nodes that have many low-distance relations in the allelic distance matrix.
Subsections
