Annotate with DIAMOND
The Annotate with DIAMOND tool allows you to annotate a DNA sequence using a set of known protein reference sequences. This tool can be used on sequences without any pre-existing annotations: it is not necessary to annotate the DNA sequences with genes or coding regions. For more information about the DIAMOND aligner, see Annotate CDS with Best DIAMOND Hit.
The tools can be used for various purposes, e.g. transferring annotations from a known reference, annotate the presence of AMR or virulence markers in a genome, or to filter contigs or sequences based on the presence of a set of genes.
For annotating DNA sequences from a set of non-coding reference sequences, the Annotate with BLAST tool may be used instead. However, the Annotate with DIAMOND tool is in general the fastest option when working with coding regions.
If the input sequences are already annotated with CDS annotations, it is also possible to use the Annotate CDS with Best BLAST Hit and Annotate CDS with Best DIAMOND Hit tools - see Annotate CDS with Best BLAST Hit for more information.
To start the tool, go to:
Tools | Microbial Genomics Module (![]() ) | Functional Analysis (
) | Functional Analysis (![]() ) | Annotate with DIAMOND (
) | Annotate with DIAMOND (![]() )
)
The first wizard step (figure 12.5), specifies the reference and search parameters.
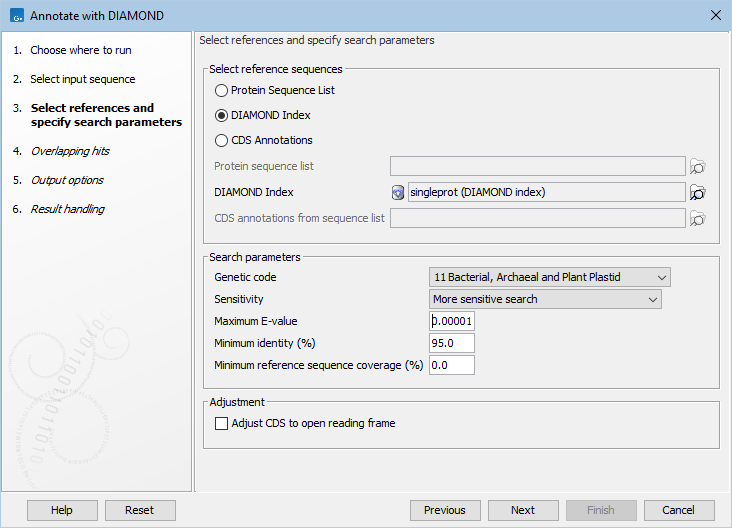
Figure 12.5: Selecting references and specifying search parameters.
The following sources can be used to annotate the input sequences:
- Protein Sequence List. The nucleotide input query will be searched against the sequences in the protein sequence list. The nucleotide input will be translated using the chosen genetic code. If the reference protein sequence list contains metadata, this metadata will be transferred to the resulting annotations on the input query sequence.
- DIAMOND Index. A DIAMOND index can be created using the Create DIAMOND Index tool. This works similar to the Protein Sequence List option, but can be faster since the index can be reused. When using the DIAMOND index, the name and description of detected reference sequences are transferred to the input query sequence.
- CDS Annotations. This option uses a nucleotide sequence source with existing annotations as a source. All CDS annotations are extracted and translated to a protein database, which is searched similar to the previous options. All qualifiers on the detected source annotations are transferred to the input query sequence.
As can be seen above, metadata (such as GO terms and taxonomy information) is handled differently depending on the database source:
- Protein sequence list. The sequence list may contain metadata, which can be inspected in the table view of the sequence list. Such metadata is transferred to the annotations created by this tool.
- DIAMOND Index. If a DIAMOND index was created from a protein sequence list containing metadata, the original metadata will be transferred to the annotations created by this tool.
- CDS annotations from sequence list. Annotations are transferred together with any metadata qualifiers the annotations contain.
The search parameters can be modified using the following settings:
- Genetic code. The code used when translating the nucleotide sequences before searching against the protein references.
- Sensitivity: Select DIAMOND sensitivity:
- Faster search: The fastest search
- Fast search: Designed for finding hits of >90% identity
- Standard search: Designed for finding hits of >60% identity
- Mid-sensitive search: More sensitive than standard search and faster than sensitive search.
- Sensitive search: Designed for finding hits of >40% identity
- More sensitive search: Designed for finding hits of >40% identity with some motif masking disabled
- Very sensitive search: Designed for finding hits of 40% identity
- Most sensitive search: The most sensitive search
- Maximum E-value. Maximum expectation value (E-value) threshold for saving hits.
- Minimum identity (%). The minimum percent amino acid identity for a hit to be accepted. Notice: when annotating with a Protein sequence list of clustered sequences such as UniRef50, this should be lowered depending on the level of clustering in the database.
- Minimum reference sequence coverage (%). The minimum length fraction of the reference sequence that must be matched. Notice: this is length fraction per hit (HSP), and should be kept low when searching for non-contiguous matches.
Adjustment can be made to the annotation hits by the following setting:
- CDS adjustment: The found annotation hits will be adjusted to begin with a start codon, end with a stop codon and not contain any stop codons in between. The adjustment can extend the annotation to up to 110 percent of the length of the reference gene and will not be shorter than 90 percent of the reference gene length. The frame of the translation may change from the original alignment.
The next step (figure 12.6), determines how to handle when multiple overlapping hits are found on the input query sequence.
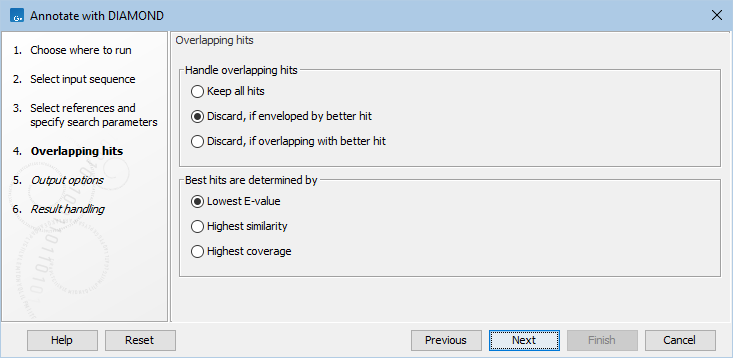
Figure 12.6: Settings for handling overlapping hits.
The following options are available:
- Keep all hits: all hits that meet the search criteria are annotated on the input query sequence.
- Discard, if enveloped by better hit: If a hit covers the same region or part of the same region as a better hit, it is discarded.
- Discard, if overlapping with better hit: If a hit overlaps the same region as a better hit, it is discarded.
Best hits are determined by:
- Lowest E-value: hits with the lowest E-value are kept. Ties are resolved by highest similarity, subsequently highest coverage.
- Highest similarity: hits with the highest similarity are kept. Ties are resolved by lowest E-value, subsequently highest coverage.
- Highest coverage: hits with the highest coverage are kept. Ties are resolved by lowest E-value, subsequently highest similarity.
The output options step (figure 12.7), has the following options:
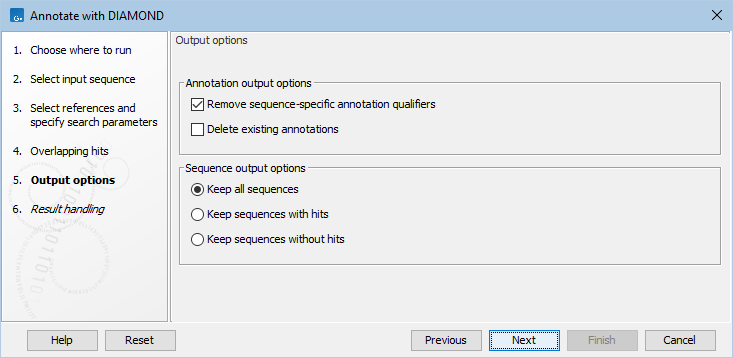
Figure 12.7: Specifying output options.
- Remove sequence-specific annotation qualifiers. Annotation qualifiers such as 'translation' and 'codon_start' may no longer be accurate on the new annotations. This option removes such qualifiers.
- Delete existing annotations. Existing annotations will not be copied to the output sequences.
The following sequence output options are available:
- Keep all sequences
- Keep sequences with hits. This option can be useful for filtering input sequences for certain regions.
- Keep sequences without hits. This option can be useful for comparing sequence lists.
The final step controls which outputs are created. Notice, that reports can be aggregated using the Combine Reports tool.
