Import RNAcentral Database
This tool can be used to import non-coding RNA sequences from RNAcentral, and join the sequences with functional Gene Ontology information.
The imported sequences can then be used together with the Annotate with BLAST tool (see section "Annotate with BLAST") and the Build Functional Profile tool (see section "Build functional profile") to quantify the functional annotation abundances.
The Import RNAcentral Database tool uses a special FASTA importer that allows for non-standard nucleotides (RNAcentral includes sequences with non-standard IUPAC nucleotide symbols, which are not allowed by our standard FASTA importer).
The tool can also import RNAcentral files with associations to GO-terms, such as 'rnacentral_rfam_annotations.tsv.gz', and match the entries with those in the imported sequence list.
Before running the tool, it is necessary to download the relevant sequences and GO-associations from RNAcentral (https://rnacentral.org/). To get the full set of annotations, we recommend downloading the following files:
RNAcentral FASTA sequences: https://ftp.ebi.ac.uk/pub/databases/RNAcentral/current_release/sequences/rnacentral_active.fasta.gz
RNAcentral GO Associations (from RFAM): https://ftp.ebi.ac.uk/pub/databases/RNAcentral/current_release/go_annotations/rnacentral_rfam_annotations.tsv.gz
To run the tool, go to:
Tools | Microbial Genomics Module (![]() ) | Databases (
) | Databases (![]() ) | Functional Analysis (
) | Functional Analysis (![]() ) | Import RNAcentral Database (
) | Import RNAcentral Database (![]() )
)
In the tool dialog (figure 17.9), select the files downloaded as described above.
It is also possible to select whether to include only RNAcentral sequences with matching GO associations, which will reduce the size of the created database.
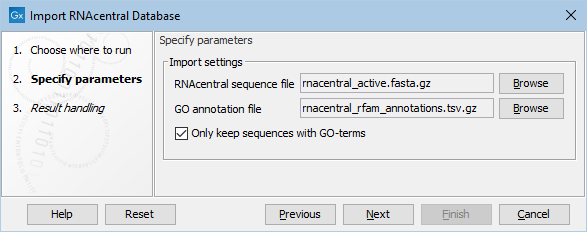
Figure 17.9: The Import RNAcentral Database tool options.
RNAcentral identifiers may contain a species-specific suffix (e.g. URS0000000006_1317357 - here 1317357 is an NCBI Taxonomy ID). When we perform the matching of RNAcentral sequences to GO associations these are stripped off and ignored.
