Identify MLST Scheme from Genomes
This section describes how to identify the relevant MLST Scheme for a genome sequence or list of genome sequences.
This tool can be used before running Type with MLST Scheme in case you are working with a sample containing a single or multiple unknown species, as in the Type among Multiple Species workflow.
Identify MLST Scheme from Genomes is available from:
Tools | Microbial Genomics Module (![]() ) | Typing and Epidemiology (
) | Typing and Epidemiology (![]() ) | MLST Typing (
) | MLST Typing (![]() ) | Identify MLST Scheme from Genomes (
) | Identify MLST Scheme from Genomes (![]() )
)
The input to the tool is a sequence, or a sequence list (figure 10.10).
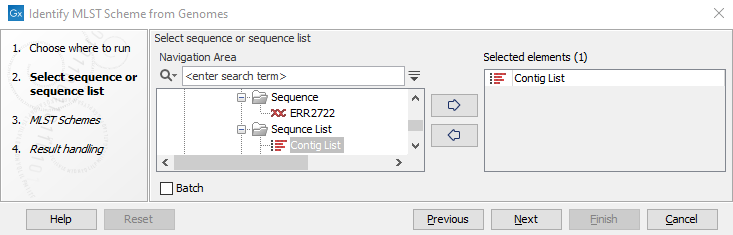
Figure 10.10: Select relevant genome sequence or sequence list.
The next step is to select as many MLST Schemes as necessary to identify the species present in the input sample (figure 10.11).
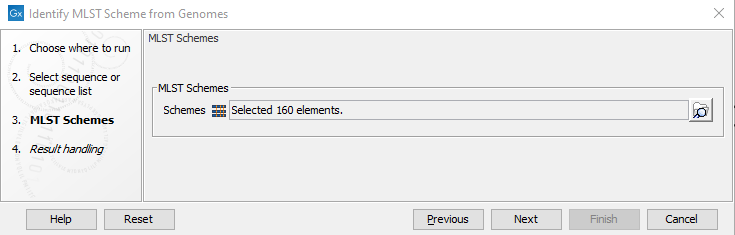
Figure 10.11: Select relevant MLST Scheme(s) to search among.
To identify the best matching scheme, the tool identifies the 10 most prevalent loci, i.e. loci that occur in most or all of the sequence types. If fewer loci are available, the tool will base the identification on these, thus the tool also works for classic 7-gene MLST schemes, given that they are in the MLST Scheme format.
The k-mers for all alleles for these most prevalent loci are then determined, and the provided references are checked for their presence.
The output of this tool is the MLST Scheme that best matches the sequences analyzed. To add the obtained best match to a Result Metadata Table, see Extend Result Metadata Table.
The tool will not produce an output if no scheme could be uniquely identified.
