Create Taxonomic Profiling Index
This tool will generate a taxonomic profiling index from a reference database. Taxonomic profiling indexes are used as input for e.g., the Taxonomic Profiling tool (Taxonomic Profiling) and the Assign Taxonomies to Sequences in Abundance Table tool (Assign Taxonomies to Sequences in Abundance Table).
The computation of index files for taxonomic profiling is memory and hard-disk intensive due to the large sizes of reference databases usually employed for this task. The algorithm requires roughly the number of bases in bytes of memory, i.e., approximately the size of the uncompressed reference database; and twice this amount in hard disk space.
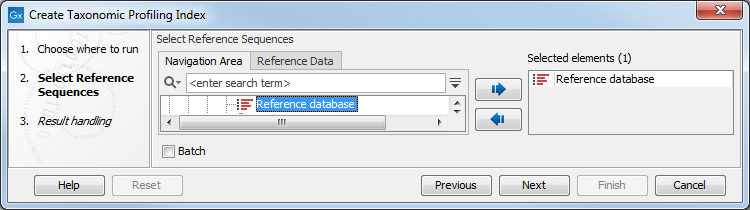
Figure 16.8: Select sequence lists with the references of interest.
To run the tool, go to:
Tools | Microbial Genomics Module (![]() ) | Databases (
) | Databases (![]() ) | Taxonomic Analysis (
) | Taxonomic Analysis (![]() ) | Create Taxonomic Profiling Index (
) | Create Taxonomic Profiling Index (![]() )
)
Select one or more sequence lists containing the references of interest. These can be downloaded for example using Download Custom Microbial Reference Database (Download Custom Microbial Reference Database).
The tool makes use of Assembly IDs (see Using the Assembly ID annotation) in combination with either Latin name or, if Latin name is not present, Sequence name. The tool will treat sequences as one reference, if they have:
- Identical Assembly ID and same Latin name, or
- Identical Assembly ID and same unique sequence name
The output is a taxonomic profiling index and an optional report as seen in figure 16.9. The report lists the number of sequence and base pairs that were indexed.
Figure 16.9: The reference sequences, index and report as seen in the Navigation Area.
