Create a Result Metadata Table
A Result Metadata Table is generated from a CLC Metadata Table with associated sample data.
To run the tool, go to:
Tools | Utility Tools (![]() ) | Result Metadata (
) | Result Metadata (![]() ) | Create Result Metadata Table (
) | Create Result Metadata Table (![]() )
)
Select the CLC Metadata Table (figure 20.4) and click on Next to specify result handling.
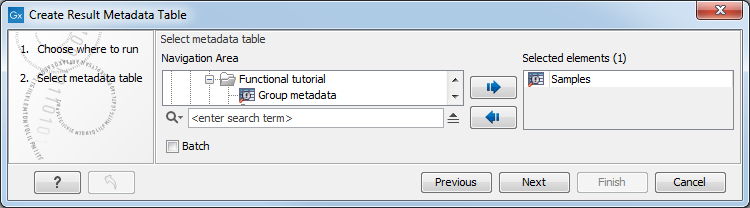
Figure 20.4: Creation of a Result Metadata Table from a Metadata Table.
The tool outputs a Result Metadata Table.
When first opened, the Result Metadata Table is empty (figure 20.5).
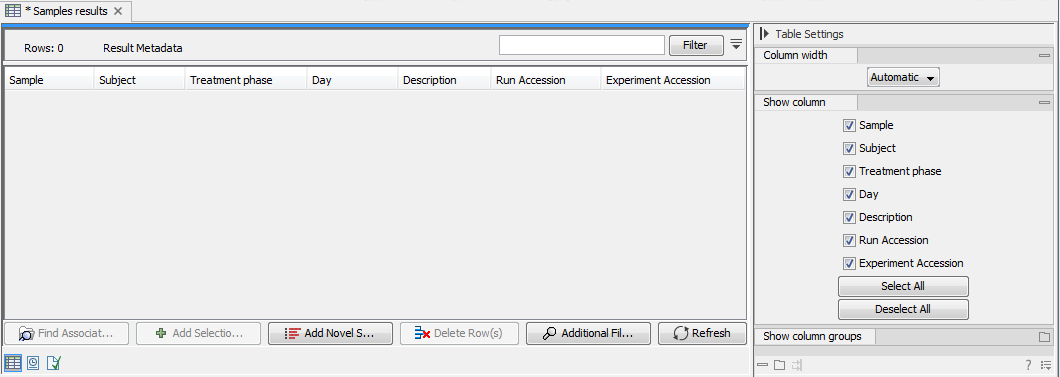
Figure 20.5: The newly created Result Metadata Table is empty.
To populate the table with information from the underlying CLC Metadata Table, click on Add Novel Samples (![]() ). Samples and associated metadata will be listed marked in yellow (figure 20.6). Save the Result Metadata Table to store the sample and metadata information.
). Samples and associated metadata will be listed marked in yellow (figure 20.6). Save the Result Metadata Table to store the sample and metadata information.
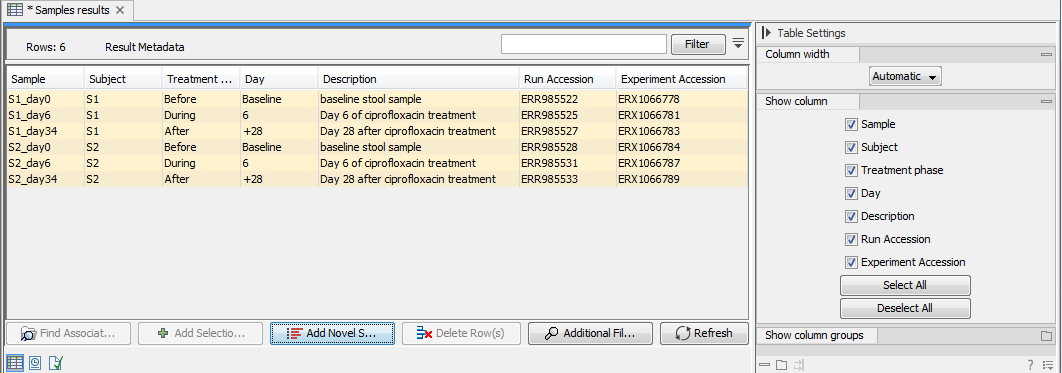
Figure 20.6: Click on "Add Novel Samples" to add metadata from the underlying CLC Metadata Table to the otherwise empty Result Metadata Table.
If for some reason Result Metadata rows are not needed, they can be deleted from the table by selecting them and clicking on the Delete Row(s) (![]() ) button.
) button.
To find files associated to specific Metadata rows, select the sample row(s) of interest and click on Find Associated Data (![]() ). This action will list all associated files in a new Metadata Element window located below the Metadata window as shown in figure 20.7.
). This action will list all associated files in a new Metadata Element window located below the Metadata window as shown in figure 20.7.
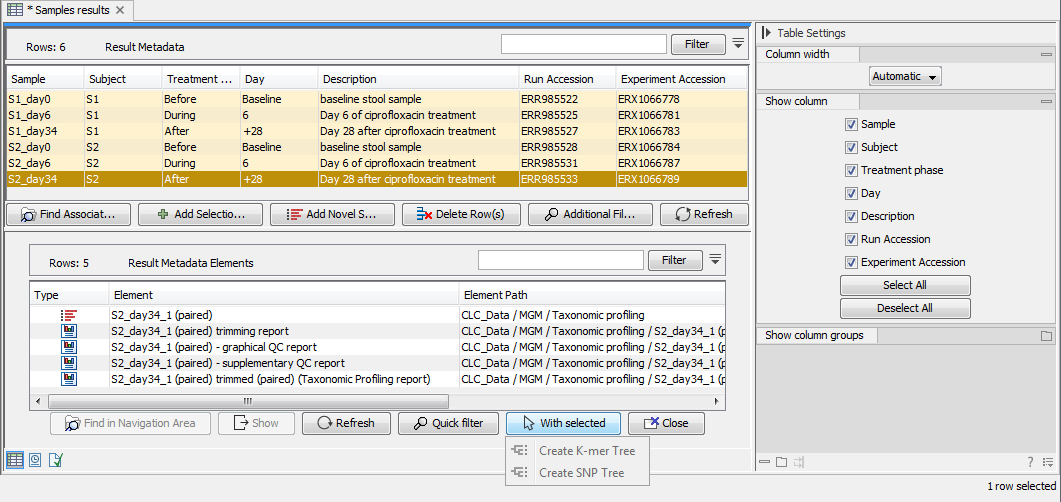
Figure 20.7: The Metadata Element window at the bottom part of this figure lists all data associated to the selected Result Metadata row shown in the top window. In this example, only the imported read file is associated to the single metadata row. Note! Workflow analysis can be initiated directly on "With selected" Elements.
In most cases, analysis results will be added automatically to the Result Metadata Table when using a properly designed workflow. It is also possible to add manually generated analysis results to the table using the Extend Result Metadata Table.
