Extracting a subset of a database
After download, it is possible to select a subset of sequences and saving the reduced list in a new sequence list. This can reduce subsequent analysis runtime significantly.
For example, from a collection of bacterial genomes that include multiple representatives of each genus, you can extract a genus specific subset of sequences to a new list:
- Open the downloaded bacterial genomes database.
- Switch to tabular element mode (
 ).
).
- Filter towards the desired genus (figure 15.2).
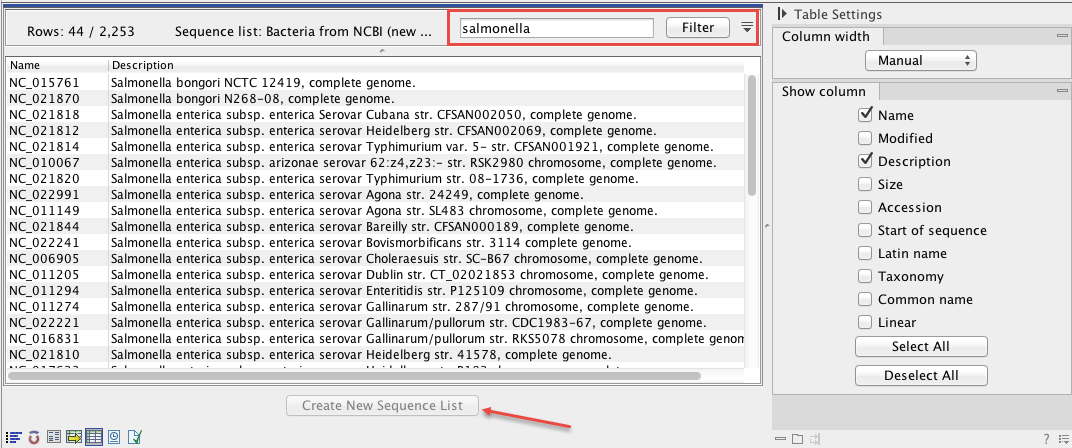
Figure 15.2: The downloaded NCBI bacterial genomes database was filtered for Salmonella data. A subset of 44 out of 2,253 sequences matched this search criterion. - Select all remaining rows.
- Click the Create New Sequence List button.
- Save the subset reference list.
Another way to extract a subset of a database is to make use of the Split Sequence List tool. For more information, see https://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=Split_Sequence_List.html.
