How to run the "Identify and Annotate Variants" ready-to-use workflow
- Go to the toolbox and double-click on the "Identify and Annotate Variants" ready-to-use workflow
(figure 14.46).

Figure 14.46: The ready-to-use workflows are found in the toolbox.This will open the wizard shown in figure 14.47 where you can select the sequencing reads from the sample that should be analyzed.
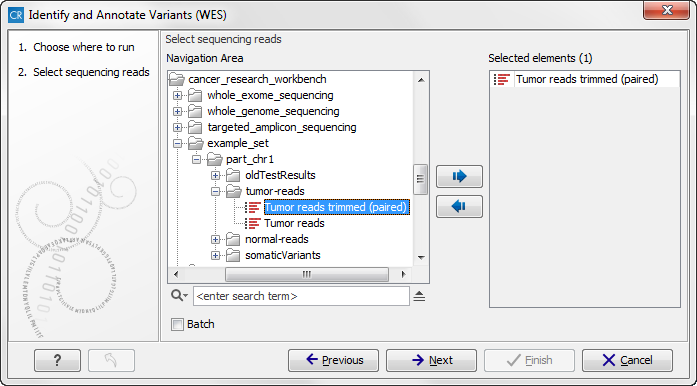
Figure 14.47: Please select all sequencing reads from the sample to be analyzed.If several samples should be analyzed, the tool has to be run in batch mode. This is done by selecting "Batch" (tick "Batch" at the bottom of the wizard as shown in figure 14.47) and select the folder that holds the data you wish to analyse. If you have your sequencing data in separate folders, you should choose to run the analysis in batch mode.
When you have selected the sample(s) you wish to prepare, click on the button labeled Next.
- In the next wizard step (figure 14.48) you can select the population from the 1000 Genomes project that you would like to use for annotation.
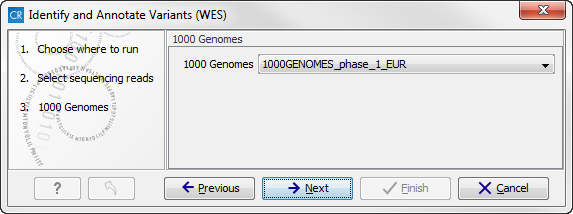
Figure 14.48: Select the population from the 1000 Genomes project that you would like to use for annotation. - In the next wizard (figure 14.49) you can select the target region track and specify the minimum read coverage that should be present in the targeted regions.
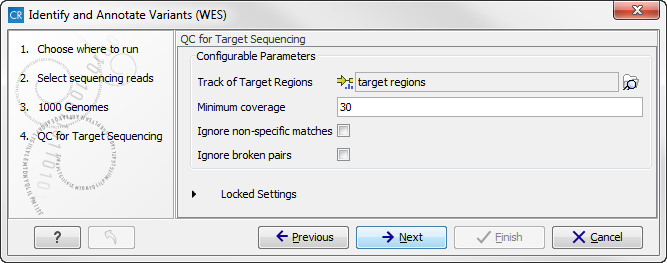
Figure 14.49: Select the track with targeted regions from your experiment. - Click on the button labeled Next, which will take you to the next wizard step (figure 14.50). In
this dialog, you have to specify the parameters for the variant detection. For a description of the different parameters that can be adjusted in the variant detection step, we refer to the description of the "Low Frequency Variant Detection" tool in the CLC Cancer Research Workbench user manual (http://www.clcsupport.com/clccancerresearchworkbench/current/index.php?manual=Low_Frequency_Variant_Detection.html). As general filters are applied to the different variant detectors that are available in CLC Cancer Research Workbench, the description of the filters are found in a separate section called "Filters" (see http://www.clcsupport.com/clccancerresearchworkbench/current/index.php?manual=Variant_Detectors_filters.html). If you click on "Locked Settings", you will be able to see all parameters used for variant detection in the ready-to-use workflow.
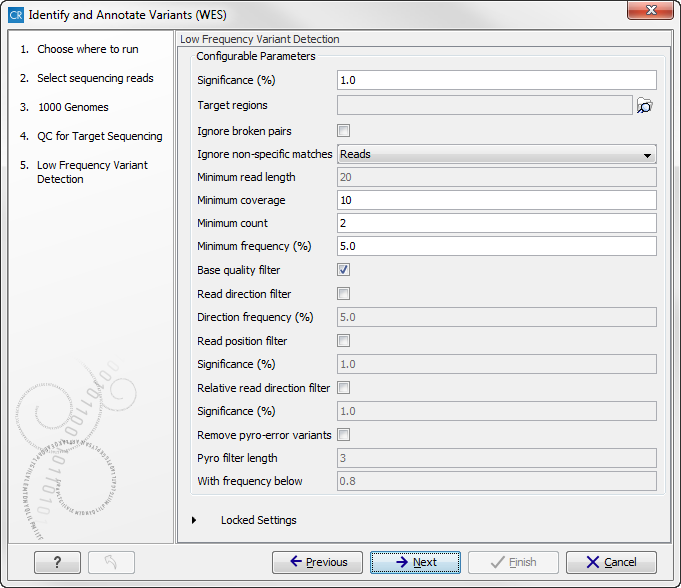
Figure 14.50: Specify the parameters for variant calling. - Click on the button labeled Next, which will take you to the next wizard step (figure 14.51). In this dialog you
can specify the target regions track. The variants found outside the targeted region will be removed at this step in the workflow.
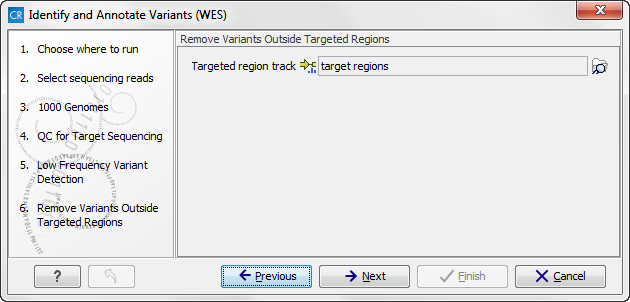
Figure 14.51: In this wizard step you can specify the target regions track. Variants found outside these regions will be removed. - Click on the button labeled Next, which will take you to the next wizard step (figure 14.52). Once again, select the relevant population from the 1000 Genomes project. This will add information from the 1000 Genomes project to your variants.
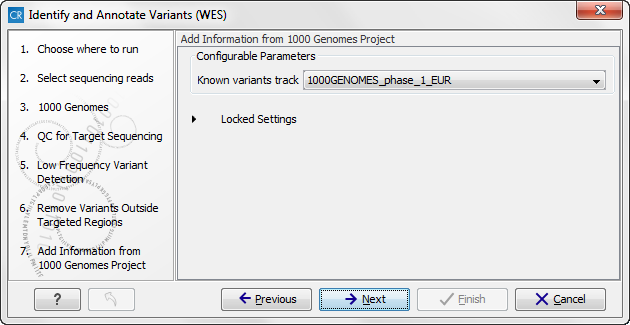
Figure 14.52: Select the relevant population from the 1000 Genomes project. This will add information from the 1000 Genomes project to your variants. - Click on the button labeled Next, which will take you to the next wizard step (figure 14.53). At this step you can select a population from the HapMap database. This will add information from the Hapmap database to your variants.
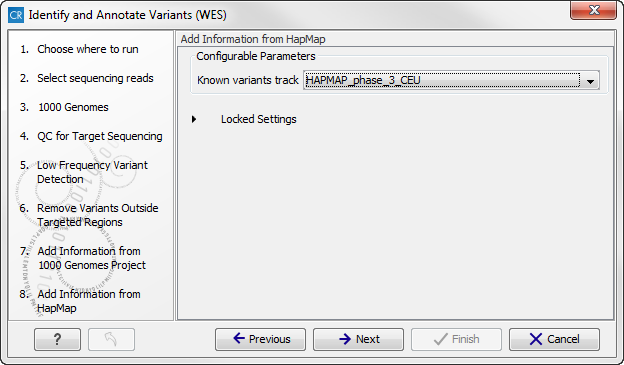
Figure 14.53: Select a population from the HapMap database. This will add information from the Hapmap database to your variants. - In this wizard step (figure 14.54) you get the chance to check the selected settings by clicking on the button labeled Preview All Parameters.
In the Preview All Parameters wizard you can only check the settings, it is not possible to make any changes at this point.
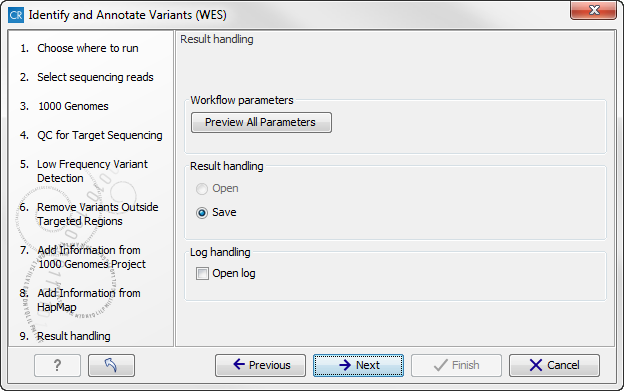
Figure 14.54: Check the settings and save your results. - Choose to Save your results and press Finish.
Note! If you choose to open the results, the results will not be saved automatically. You can always save the results at a later point.
