How to run the "Identify Known Variants in One Sample" ready-to-use workflow
- Go to the toolbox and double-click on the "Identify Known Variants from
One Sample" ready-to-use workflow
(figure 13.30).

Figure 13.30: The ready-to-use workflows are found in the toolbox.This will open the wizard step shown in figure 13.31 where you can select the reads of the sample, which should be tested for presence or absence of your known variants.
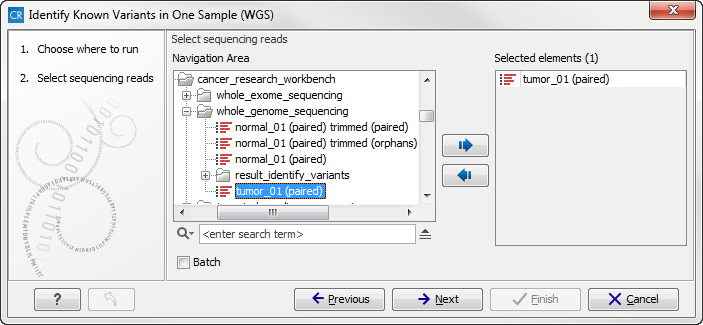
Figure 13.31: Select the sequencing reads from the sample you would like to test for your known variants.Please select all sequencing reads from your sample. If several samples should be analyzed, the tool has to be run in batch mode. This is done by selecting "Batch" (tick "Batch" at the bottom of the wizard as shown in figure 13.31) and select the folder that holds the data you wish to analyse. If you have your sequencing data in separate folders, you should choose to run the analysis in batch mode.
When you have selected the sample(s) you wish to analyze, click on the button labeled Next.
In the next wizard step, you can select your known variants that you would like to check for in your read mapping (figure 13.32).
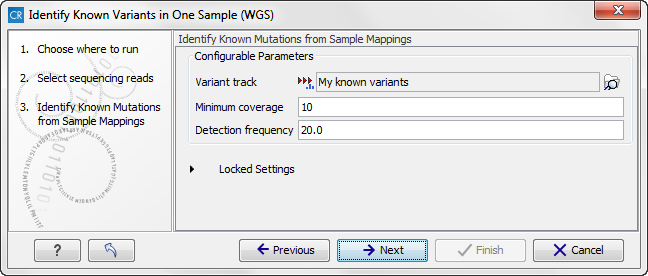
Figure 13.32: Specify the track with the known variants that should be identified.The parameter "Detection Frequency" will be used in the calculation twice. First, it will report in the result if a variant has been detected (observed frequency > specified frequency) or not (observed frequency > specified frequency). Moreover, it will determine if a variant should be labeled as heterozygote (frequency of another allele identified at a position of a variant in the alignment > specified frequency) or homozygote (frequency of all other alleles identified at a position of a variant in the alignment < specified frequency).
- Click on the button labeled Next, which will take you to the next wizard step (figure 13.33). In
this and the next dialog, you will be asked about which of the annotations/informations added to variants should be included in the results.
Please specify your track with known variants.
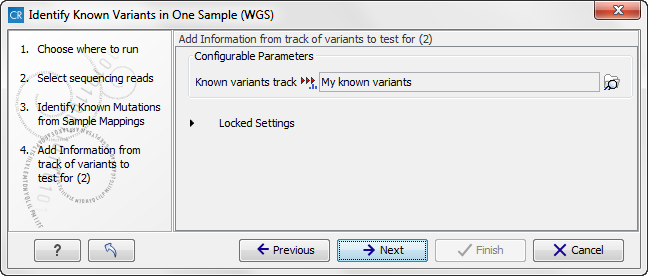
Figure 13.33: Select the track with your known variants again. Annotations/Information from this track will be added to the overview mutation track. - Click on the button labeled Next and once again select the same track with known variants (figure 13.34).
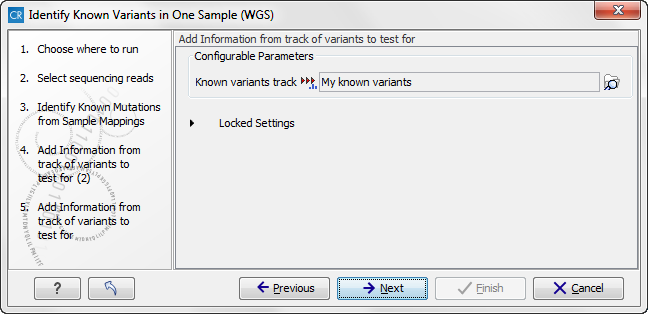
Figure 13.34: Once again select the track with known variants. This time the track is used to add information to the detailed mutation track. - Click on the button labeled Next to go to the last wizard step (figure 13.35).
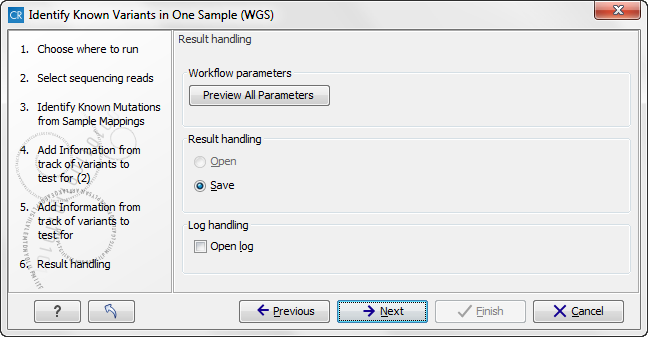
Figure 13.35: Check the settings and save your results.In this wizard step you can check the selected settings by clicking on the button labeled Preview All Parameters. In the Preview All Parameters wizard you can only check the settings, it is not possible to make any changes at this point. At the bottom of this wizard there are two buttons regarding export functions; one button allows specification of the export format, and the other button (the one labeled "Export Parameters") allows specification of the export destination. When selecting an export location, you will export the analysis parameter settings that were specified for this specific experiment.
- Click on the button labeled OK to go back to the previous dialog box and choose Save.
Note! If you choose to open the results, the results will not be saved automatically. You can always save the results at a later point.
