Prepare sequencing data - all application types
The first thing to do after data import is to check the quality of the sequencing reads and perform the necessary trimming. This applies no matter whether you are working with Whole Genome Sequencing, Exome Sequencing, or Targeted Amplicon Sequencing. In the toolbox you can choose between the two different ready-to-use workflows for data preparation that are shown in the "Run workflow 1" box in figure 12.1.The "Preparing Raw Data" ready-to-use workflows are universal and can be used for all applications; Whole Genome Sequencing, Exome Sequencing, and Targeted Amplicon Sequencing.
Choosing between "Prepare Raw Data" and "Prepare Overlapping Raw Data" workflows:
Many whole genome sequencing, exome sequencing using capture technology, and targeted amplicon sequencing strategies produce overlapping reads. Downstream stages of the Cancer Research Workbench (e.g. Variant calling) take the frequencies of observed alleles into consideration as well as the forward-reverse strand balance. When merging overlapping reads these two parameters will be affected: 1) the frequency of observed alleles in overlapping regions will be corrected (a variant found both on the forward and the reverse read of the same fragment should only be counted once), and 2) in the merged fragments the information on forward-reverse strand origin has become meaningless. These effects have to be taken into consideration when filtering variants on these statistics. As the forward-reverse strand balance statistic is used as a variant filter (i.e. the Read direction filter), we recommend using the "Prepare Overlapping Raw Data" workflow on targeted amplicon sequencing data with overlapping read sequencing strategy, whereas we recommend the "Prepare Raw Data" workflow for other sequencing protocols (e.g. whole genome sequencing, whole exome-sequencing, also if making use of overlapping read sequencing).
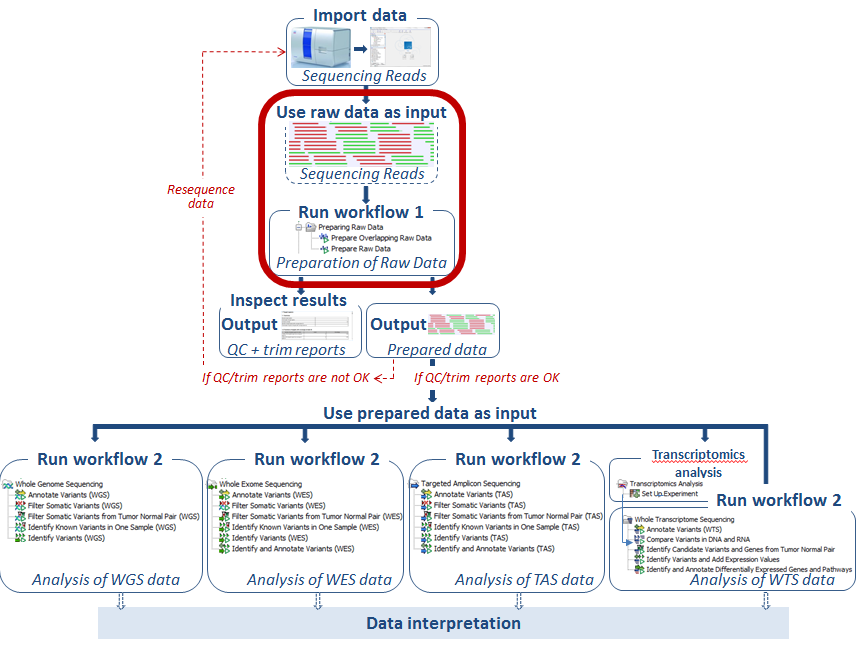
Figure 12.1: Two ready-to-use workflows are available for data preparation; "Prepare Overlapping Raw Data" and "Prepare Raw data".
Subsections
- Import adapter trim list
- How to run the "Prepare Overlapping Raw Data" ready-to-use workflow
- How to run the "Prepare Raw Data" ready-to-use workflow
- Output from the Prepare Overlapping Raw Data and Prepare Raw Data workflows
- How to check the output reports
