Finding annotations on the genome
In the Side Panel under Find, a search field allows you to quickly find the annotation that you are looking for. The list of tracks further allows you to restrict the search to a particular track (e.g. a gene track).
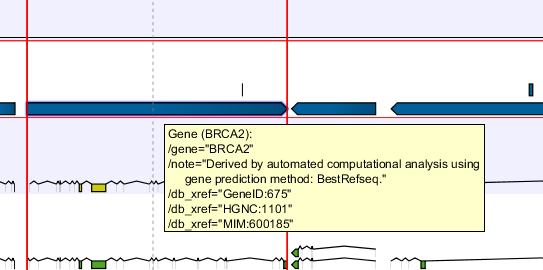
In the search field you can enter any kind of text that exists in the annotation track. As an example, consider the gene and tool tip shown in figure 18.17.
This gene could be found by searching for the name specifically or, by using a wildcards (asterisks), less specific search terms could be use, giving more flexibility. To find BRCA2, for example, any of the following entries could be typed in the search field:
- BRCA2
- This would match the annotation name exactly.
- BRCA*
- This would match the annotation name as well as other genes with a text starting with BRCA (e.g. the BRCA1 gene).
- *RCA2
- This would match the annotation name as well as other genes with a text ending with RCA2 (e.g. the SMARCA2 gene).
- 600185
- This would match the
db_xrefqualifier for the OMIM database. All the text shown for the annotation in figure 18.17 can be searched this way, both as exact matches and with the * before or after the search term.
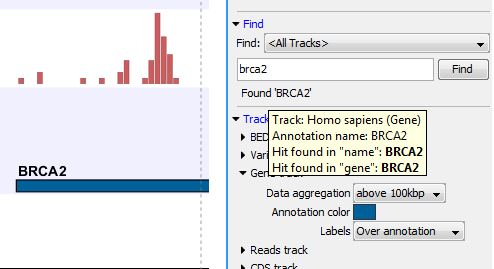
Figure 18.18: The BRCA2 gene found.
The search will be performed throughout the entire genome beginning with the chromosome currently shown and stopping when it finds the first hit. Press Find again to find the next hit. Once the whole genome has been traversed, the status will inform you that you have searched the whole genome. Click the Find button to start the search again.
Please note that you can also use the table view of an annotation track to perform more advanced queries of the data (see Showing a track in a table).