Create Filter Criteria
To run the Create Filter Criteria, go to the toolbox:
Toolbox | Identify Candidate Variants (![]() ) | Create Filter Criteria (
) | Create Filter Criteria (![]() )
)
After the variants have been identified and post-filtered (e.g. for somatic variants), the next task is to identify e.g. driver mutations or at to identify those variants, which should be validated first.
The Create Filter Criteria tool can help setting up advanced filter criteria that can be used in the Identify Candidate Variants tool (see Identify Candidate Variants) to identify candidate variants. The Identify Candidate Variants) can be used in analysis workflows later on.
An example of a filter criterion is shown in figure 26.1 where you will filter out everything that is not non-synonymous variants or variants that are present in the COSMIC database. As a result you will only keep the variants that are already known in cancer (present in the COSMIC database) or that are non-synonymous.
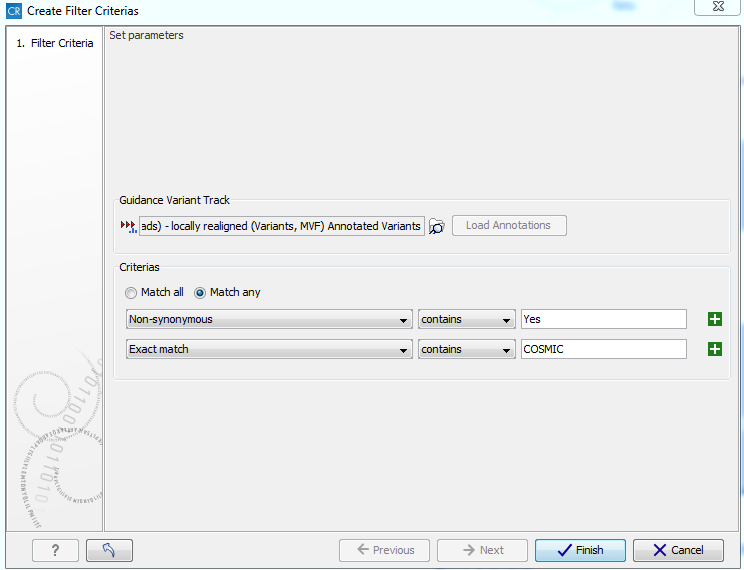
Figure 26.1: A filter criterion to extract those variants, which
are known in cancer (present in the COSMIC database) or are non-synonymous
Please select and load one of your annotated variant tracks in the dialog box by clicking on the folder icon. This track is used as a guidance variant track in order to obtain the annotation categories (annotation column headers) to be used for selection of which annotations the filter should be applied to. Click on the "Load Annotations" button to activate the filter categories found under "Criteria".
The created filter criteria can be modified by opening the generated output from the Create Filter Criteria by clicking on the name of the filter criteria file where you saved it the Navigation Area.
Caution! Please note, that when you create filter criteria, you should use a guiding variant track that contains the same annotations as the annotations that are present in the variant tracks on which the filter criteria should be applied to in a workflow.
