Running Velvet from the Workbench
To run Velvet, open a Workbench and connect to a server where the Velvet external application is installed. Then:
- Go to:
Tools | External Applications (
 ) | CLC bio Velvet (
) | CLC bio Velvet ( )
)
- Confirm where you wish to run the job.
- Select (
 ) the reads to assemble and configure the Velvet parameters, as shown in figure 16.26.
) the reads to assemble and configure the Velvet parameters, as shown in figure 16.26.
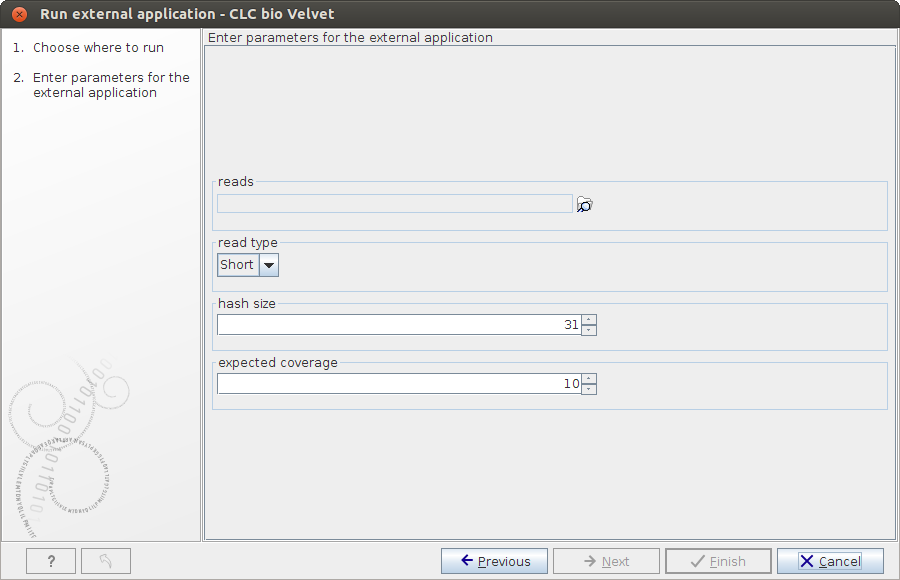
Figure 16.26: Configuring Velvet parameters from a Workbench. - Click Next.
- Specify where to save the results.
- Click Finish.
The process that follows has four steps:
- The sequencing reads are exported by the server to a fasta format file. This is a temporary file that will be deleted when the process is done.
- The velvet script is executed using the fasta file and the user-specified parameters as input.
- The resulting output file is imported into the save location specified in the save step of the Workbench dialog, and the user is notified that the process is done.
- All temporary files are deleted
