Perform QIAseq Multimodal Analysis with TMB and MSI
The Perform QIAseq Multimodal Analysis TMB and MSI (Illumina) template workflow is a copy of the Perform QIAseq Multimodal Analysis (Illumina) template workflow, extended with the functionality to calculate a TMB score as well as assess MSI status. The panel related to the workflow is the QIAseq Multimodal Pan Cancer panel (UHS-5000Z).
|
Note that the QIAseq Multimodal Panels are designed against genome build hg19 for the DNA panel and hg38 for the RNA panel. BED files are provided in the respective genome build. However, the template workflow requires that reference data for both DNA and RNA is for the same genome build. The two QIAseq Multimodal Reference Data Sets provided by the Reference Data Manager are for genome build hg38, where the reference data for the DNA panel has been converted to hg38 as described below.
For custom panels, the DNA panel BED file needs to be imported against hg19, after which it should be converted to hg38 using the tool Convert Annotation Track Coordinates. If many regions are lost during conversion, it can cause reads to be discarded that would have otherwise mapped to the lost target regions. To avoid such issues, a copy of the template workflow can be used, containing only the analysis of the DNA reads, and the workflow should be run using the imported BED file against hg19. |
Note: A default MSI baseline from the Reference Data Set is provided for this workflow, but this is for demo purpose only and will not give the true microsatellite instability status. We recommend that the MSI baseline is generated using samples that are sequenced under the same lab conditions as the multimodal samples (see Generate MSI Baseline).
The Perform QIAseq Multimodal Analysis TMB and MSI (Illumina) template workflow can be found at:
Template Workflows | Biomedical Workflows (![]() ) | QIAseq Sample Analysis (
) | QIAseq Sample Analysis (![]() ) | Other QIAseq workflows (
) | Other QIAseq workflows (![]() ) | Perform QIAseq Multimodal Analysis TMB and MSI (Illumina) (
) | Perform QIAseq Multimodal Analysis TMB and MSI (Illumina) (![]() )
)
Double-click on Perform QIAseq Multimodal Analysis TMB and MSI (Illumina) and start an analysis.
If you are connected to a CLC Server via the CLC Workbench, you will be asked where you would like to run the analysis. We recommend that you run the analysis on a CLC Server when possible.
First, specify the DNA, and in the next dialog, the RNA sequencing reads that should be analyzed (figures 15.14 and 15.15). To run the workflow with multiple samples, see Running multimodal workflows in batch using metadata.
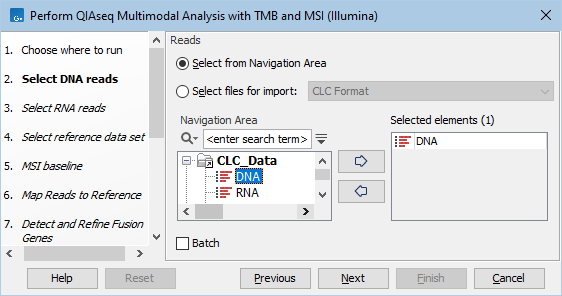
Figure 15.14: Select the sequencing reads by double-clicking on the file name or by clicking once on the file name and then on the arrow pointing to the right hand side.
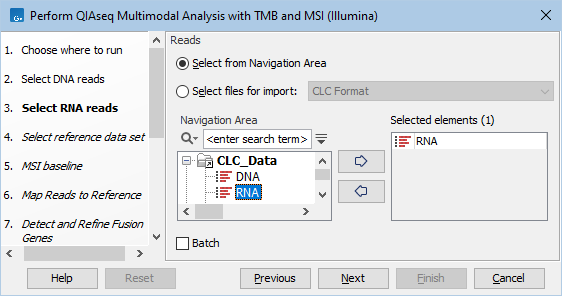
Figure 15.15: Select the sequencing reads by double-clicking on the file name or by clicking once on the file name and then on the arrow pointing to the right hand side.
The following dialog helps you set up the relevant Reference Data Set. If you have not downloaded the Reference Data Set yet, the dialog will suggest the relevant data set and offer the opportunity to download it using the Download to Workbench button. (figure 15.16).
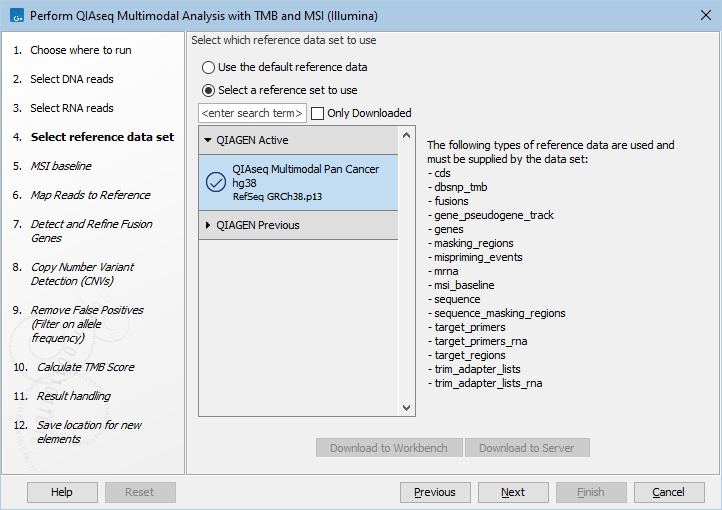
Figure 15.16: The relevant Reference Data Set is highlighted; in the text to the right, the types of reference needed by the workflow are listed.
Note that if you wish to Cancel or Resume the Download, you can close the template workflow and open the Reference Data Manager where the Cancel, Pause and Resume buttons are available.
If the Reference Data Set was previously downloaded, the option "Use the default reference data" is available and will ensure the relevant data set is used. You can always check the "Select a reference set to use" option to be able to specify another Reference Data Set than the one suggested.
In the next dialog, you are asked to select the MSI baseline. The workflow is pre-configured with a baseline from the Reference Data Set, but it is recommended to generate and use a baseline trained on samples that are sequenced under the same lab conditions as the multimodal samples you are analyzing (see Generate MSI Baseline).
In the Map Reads to Reference dialog, it is possible to configure masking. A custom masking track can be used, but by default, the masking track is set to GenomeReferenceConsortium_masking_hg38_no_alt_analysis_set, containing the regions defined by the Genome Reference Consortium, which serve primarily to remove false duplications, including one affecting the gene U2AF1. Changing the masking mode from "No masking" to "Exclude annotated" excludes these regions.
In the Detect and Refine Fusion Genes dialog, it is possible to change the Promiscuity threshold, i.e., the maximum number of different fusion partners reported for a gene. You can also check for exon skippings by enabling the "Detect exon skippings" option, as well as check for fusions with novel exon boundaries by enabeling the "Detect fusions with novel exon boundaries" option. This dialog is shown in figure 15.17
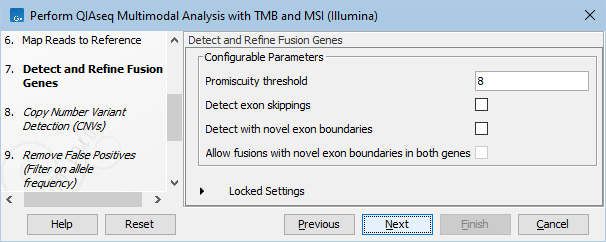
Figure 15.17: The Detect and Refine Fusion Genes dialog.
The QC for Target Sequencing cannot be modified in this workflow and the coverage restriction is kept at 100x for calculating TMB score.
The dialog for Copy Number Variant Detection allows you to specify a control mapping against which the coverage pattern in your sample will be compared in order to call CNVs. If you do not specify a control mapping, or if the target regions files contains fewer than 50 regions, the Copy Number Variation analysis will not be carried out.
Please note that if you want the copy number variation analysis to be done, it is important that the control mapping supplied is a meaningful control for the sample being analyzed. Mapping of control samples for the CNV analysis can be done using the workflows described in Create QIAseq DNA CNV Control Mapping workflows.
A meaningful control must satisfy two conditions: (1) It must have a copy number status that it is meaningful for you to compare your sample against. For panels with targets on the X and Y chromosomes, the control and sample should be matched for gender. (2) The control read mapping must result from the same type of processing that will be applied to the sample. One way to achieve this is to process the control using the workflow (without providing a control mapping for the CNV detection component) and then to use the resulting UMI reads track as the control in subsequent workflow runs.
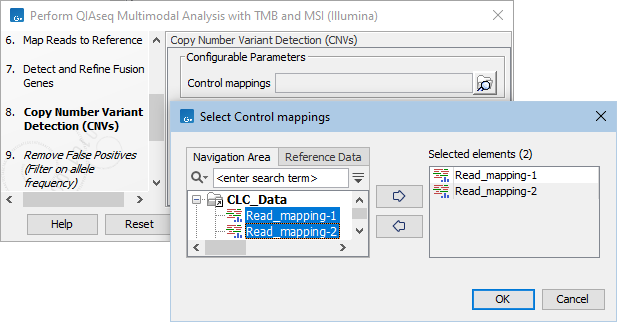
Figure 15.18: The Copy Number Variation Detection dialog. Here 3 control samples have been selected. In practice it is recommended to either use a matched control sample, or to use at least 5 control samples. Increasing the number of samples beyond this does not typically improve results.
The parameters for variant detection are not adjustable and have been set to generate an initial pool of all potential variants. These are then passed through a series of filters to remove variants that are suspected artifacts. Variants failing to meet the (adjustable) thresholds for quality, read direction bias, location (low frequency indels within homopolymer stretches), frequency or coverage would not be included in the filtered output.
Some filters only remove alternative alleles - and not reference alleles - as this potentially leads to wrong interpretation of variants by the VCF exporter where such variants could be misinterpreted as hemizygote when the reference allele is missing.
The frequency cutoff is the only open parameter in this workflow and the workflow can detect down to 1% variant frequency. Even when setting the frequency lower it will not output lower frequencies as the variant calling is initially done down to 1% by the variant caller in this workflow. Further adjustments needs to be done by opening a copy of the workflow.
Configure the result handling and in the last dialog, choose to Save the results of the workflow and specify a location in the Navigation Area.
Subsections
