Output for the Perform QIAseq FastSelect RNA Expression and Fusion Calling (N6-T RT primer) workflow
The Perform QIAseq FastSelect RNA Expression and Fusion Calling (N6-T RT primer) workflow produces outputs as shown in figure 14.9. The figure shows the results of using the workflow on two samples.
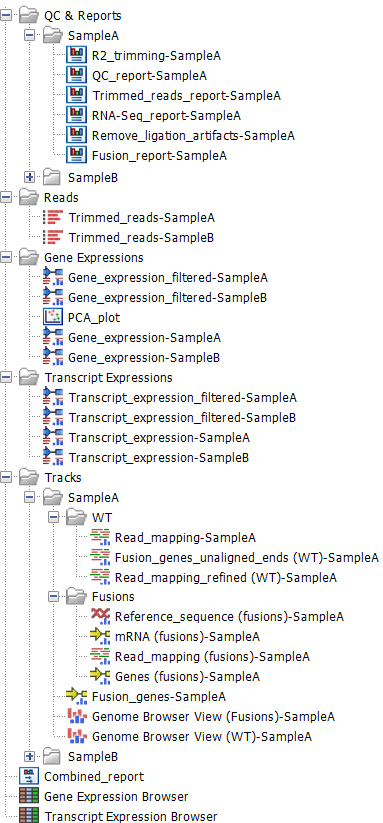
Figure 14.9: Output from the Perform QIAseq FastSelect RNA Expression and Fusion Calling (N6-T RT primer) workflow when run with two samples.
Two Expression Browsers (![]() ) tabulate the transcript level expressions of all the samples and the gene level expressions of all the samples.
) tabulate the transcript level expressions of all the samples and the gene level expressions of all the samples.
The Combined_report (![]() ) summarizes all the reports generated for all the samples during analysis.
) summarizes all the reports generated for all the samples during analysis.
The QC & Reports folder provides detailed reports (![]() ) for individual tools run in the workflow. There is one subfolder per sample. Reports of special interest are:
) for individual tools run in the workflow. There is one subfolder per sample. Reports of special interest are:
- RNA-Seq_report A graphical report of the RNA-seq analysis. See https://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=RNA_Seq_report.html.
- Fusion_report A graphical report of the fusions found in the sample. For more details see https://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=Output_from_Detect_Refine_Fusion_Genes_tool.html.
The Reads folder contains the trimmed reads used in the downstream analysis.
The Gene Expressions and Transcript Expressions folders contain two types of Expression Track (![]() ) - one at the gene level and one at the transcript level. A "filtered" copy of these tracks is also provided. The filtered tracks are limited to genes/transcripts that are annotated as being "protein coding" or "lncRNA". Further details about this track type can be found at https://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=Expression_tracks.html.
) - one at the gene level and one at the transcript level. A "filtered" copy of these tracks is also provided. The filtered tracks are limited to genes/transcripts that are annotated as being "protein coding" or "lncRNA". Further details about this track type can be found at https://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=Expression_tracks.html.
Within the Gene Expressions folder is a PCA plot. This will be empty if there is only a single sample. When there are multiple samples, the clustering in the PCA plot can be used for quality control. For more details on PCA plots see https://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=PCA_RNA_Seq.html.
The Tracks folders contain the results of fusion calling. There is one subfolder per sample. The results are split into two parts - the "WT" (wildtype) chromosomes, showing reads that align to the reference genome, and the "Fusions" chromosomes showing reads that preferentially align to the fusion products. All the data for the wildtype and fusion chromosomes can be viewed by opening the "Genome Browser View (WT)" (![]() ) and "Genome Browser View (Fusions)" (
) and "Genome Browser View (Fusions)" (![]() ) respectively. These show the data in the "WT" and "Fusions" folders.
) respectively. These show the data in the "WT" and "Fusions" folders.
For quality control of fusion calls, see https://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=Interpretation_fusion_results.html. We particularly recommend carrying out manual quality control checks on results that include fusions with novel exon boundaries.
