Sankey plot for Clonotype Sample Comparison
The Sankey plot (
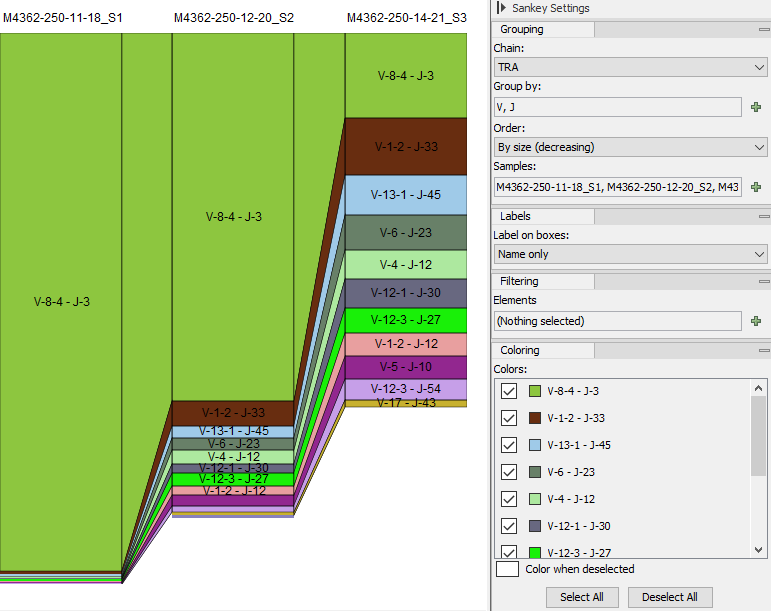
Figure 7.26: Sankey plot for the TRA chain showing frequencies of clonotypes with specific V and J segments, compared across samples.
For each selected sample, the plot has a column that contains boxes for each group of clonotypes (hereby referred to as simply clonotypes) with the selected properties. The properties, such as the segment type or the CDR3 amino acid sequence, are selected from the side panel under "Group by".
The height of a box indicates the frequencies of the clonotypes in the sample. The frequency is used instead of the count to make samples with a different number of reads comparable.
Note that the sum of the height of boxes may differ across samples. The frequency, across all chains, adds up to 100% when clonotypes are first constructed. It is unlikely that the sum of the frequencies for a specific chain adds up to the same total for two different samples. To achieve this, the clonotypes can be filtered to only contain the desired chain and the frequencies can be recalculated. Additionally, filtering can lead to frequencies adding to less than 100%. See Filter Immune Repertoire for details on how to filter.
