Calculate Unique Molecular Index Groups
The Calculate Unique Molecular Index Groups tool annotates the mapped reads with a "Unique Molecular Index group ID", that is identical for reads that are determined to belong to the same UMI.
Calculate Unique Molecular Index Groups is available under the Tools menu at:
Tools | Biomedical Genomics Analysis (![]() ) | UMI Tools (
) | UMI Tools (![]() ) | Calculate Unique Molecular Index Groups (
) | Calculate Unique Molecular Index Groups (![]() )
)
In the first dialog (figure 4.2), select a read mapping of reads that were previously annotated with UMI annotations.
Figure 4.2: Select a read mapping made from reads whose UMI was removed and annotated on the sequences.
The grouping of reads into UMI groups works as follows:
- The tool groups reads that
- start at the same position based on the end of the read to which the UMI is ligated. This can either be defined in the Remove and Annotate with Unique Molecular Index tool or directly in the wizard, (If the UMI was removed from the start of read 2 using the Remove and Annotate with Unique Molecular Index tool, this tool considers grouping reads where the start of read 2 map to the same position)
- are from the same strand, and
- have identical UMIs.
- Their start positions are sufficiently close as defined by the Window size parameter.
- Their UMIs are similar enough as defined by the Fuzzy match Unique Molecular Indices parameter.
Merging is only done if the larger group is sufficiently large compared to the smaller group as defined by the parameters described below. If a smaller group can be merged into multiple larger groups that are equally good in terms of similarity of UMI and start position as well as group size, the group will not be merged.
Duplex groups are created if input reads were defined as duplex data in the Remove and Annotate with Unique Molecular Index tool, or if the UMI location setting has been set to duplex. Duplex groups consist of two paired end UMI groups from different strands of the original fragment. Two UMI groups, A and B, are grouped to a duplex group when:
- Both are paired reads.
- One must consist of forward paired end reads (referred to as group A) and the other must consist of reverse paired end reads (referred to as group B)
- The genomic positions of read 1 in group A and read 2 in group B are the same, and the genomic positions of read 2 in group A and read 1 in group B are the same.
- The UMI is the same for read 1 of group A and read 2 of group B, and the UMI is the same for read 2 of group A and read 1 of group B.
It is possible to change the following parameters (figure 4.3):
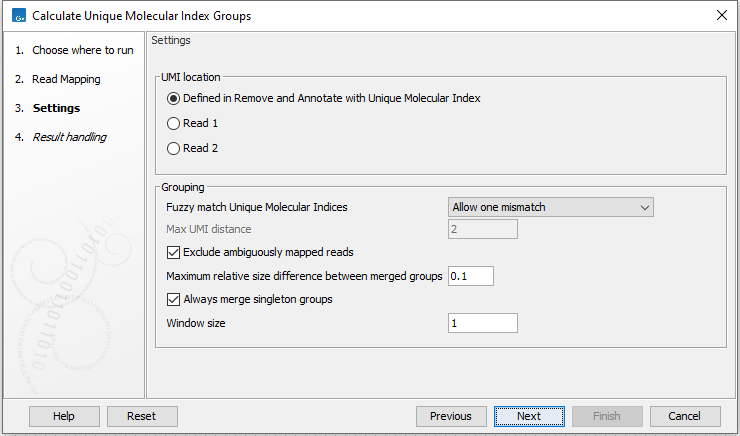
Figure 4.3: Select a read mapping made from reads whose UMI was removed and annotated on the sequences.
- UMI location Define if the UMI was sequenced as part of read 1 or read 2. The UMI location determines if the R1 or R2 start position must be the same for reads to be grouped.
- Defined in Remove and Annotate with Unique Molecular Index Groups reads based on the mapped genomic position of the start of the read that originally had the UMI as defined in Remove and Annotate with Unique Molecular Index.
- Read 1 Groups reads based on the mapped genomic position of the start of read 1. Use if the UMI sequence was annotated on the reads during import, and the UMI was originally sequenced as part of read 1. Read about annotating with UMIs during import here: https://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=General_notes_on_UMIs.html
- Read 2 Groups reads based on the mapped genomic position of the start of read 2. Use if the UMI sequence was annotated on the reads during import, and the UMI was originally sequenced as part of read 2. Read about annotating with UMIs during import here: https://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=General_notes_on_UMIs.html
- Read 1 and read 2 Groups reads based on the mapped genomic positions of the start of read 1 and read 2. Does not calculate duplex groups.
- Read 1 and read 2 (duplex) Groups reads based on the mapped genomic positions of the start of read 1 and read 2. And also calculates duplex groups.
- Fuzzy match Unique Molecular Indices Method for deciding which UMIs are considered similar enough for merging:
- Do not fuzzy match Groups will be merged only if they match exactly.
- Allow one mismatch Groups will be merged if they are at most one mismatch apart.
- Allow one mismatch/deletion/insertion Groups will be merged if they are at most one mismatch, deletion or insertion apart.
- Distance Groups will be merged if their edit distance (also called Levenshtein distance), is smaller than the value given in Max UMI distance. Note that if the distance is greater than 1, the groups also have to satisfy a stricter requirement for ratio between their sizes.
- Max UMI distance The maximum edit distance allowed if Distance is selected as Fuzzy match Unique Molecular Indices.
- Exclude ambiguously mapped reads is checked by default.
- Maximum relative size difference between merged groups will merge small groups into bigger ones if the size ratio between the two groups is smaller than a certain value (set at 0.1 per default).
We define the distance between two groups as the number of differences in their UMIs (which can only be greater than one if "Distance" is chosen as Fuzzy match Unique Molecular Indices) plus one if their start positions are not the same.
The size ratio parameter is taken to the power of the distance, i.e. if the distance is two, the smaller group size should be at most
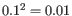 the size of the larger group.
the size of the larger group.
- Always merge singleton groups When this option is checked, a singleton UMI group, a group that contain only one read, is merged with a non-singleton group with distance 1 even if the "Maximum relative size difference between merged groups" threshold is not met.
- Window size Groups will be merged if the difference between their start positions are less than this.
Click Next to choose whether to Open or Save the resulting read mapping of reads which now have a "UMI group ID" annotation.
A report can also be generated. It contains:
- A summary table with the following information:
- Reads in input: Reads that were aligned to the reference
- Reads mapped multiple places (discarded): Reads that aligned to the reference in multiple places, and thus discarded
- Groups merged
- Groups not merged due to
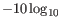 1 candidate of equal size
1 candidate of equal size
- Group size table and plots described in UMI group sizes.
- Duplex summary and plots if calculating duplex groups.
Note: When the group sizes (the number of reads in UMI groups) are very large (in most cases more than 10 reads in a UMI group is not desirable), this can indicate problems, such as quality issues with the sample. It can also indicate that the sequencing depth could be reduced.
