Import Known Fusion Information Track
Import Known Fusion Information Track can import information about known fusions from a tab separated text file. It outputs an annotation track (![]() ) that can be used with Annotate Fusions with Known Fusion Information.
) that can be used with Annotate Fusions with Known Fusion Information.
The importer can be found here:
Import (![]() ) | Import Known Fusion Information Track (
) | Import Known Fusion Information Track (![]() )
)
The following options can be configured (figure 5.6):
- File A tab separated text file with the known fusion information. The first row in the file is a header. Each subsequent row lists the information for a known fusion.
Information is extracted from the following columns in the file:
5prime_geneand3prime_gene(mandatory): the name of the 5'/3' gene.5prime_bpand3prime_bp(optional): the position of the 5'/3' breakpoint.The position can be exact (e.g., 1364892), or uncertain (e.g., 1364887-1364895).
5prime_chrand3prime_chr(optional): the chromosome of the 5'/3' gene.5prime_strandand3prime_strand(optional): the strand (+/-) of the 5'/3' gene.
Further columns are added as additional attributes.
- Gene track An annotation track (
 ) with the gene annotations. The 5' and 3' genes in the provided file are matched to this track by gene names, with support for synonyms. If chromosome and/or strand information is available in the file:
) with the gene annotations. The 5' and 3' genes in the provided file are matched to this track by gene names, with support for synonyms. If chromosome and/or strand information is available in the file:
- For genes that match the track:
- If the chromosome in the file and track do not match, the fusion is not imported.
- If the strand in the file and track do not match, the strand from the track is used.
- If the breakpoint position is not available, the entire gene region is used.
- For genes that do not match the track:
- The fusion is imported with the chromosome/strand provided in the file.
- Fusions with genes without a breakpoint position are not imported.
- For genes that match the track:
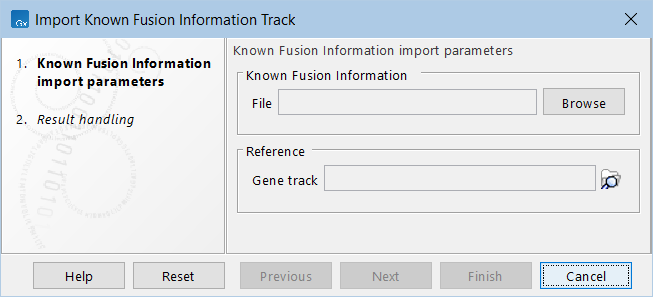
Figure 5.6: The available options when importing known fusion information.
