Output from the Quantify QIAseq RNA Expression workflow
The Quantify QIAseq RNA Expression template workflow produces the following outputs:
- Expression tracks (
 ) for each sample.
) for each sample.
- Sample report (
 ): A report containing essential information from all reports produced by the workflow. See Create Sample Report output for details.
): A report containing essential information from all reports produced by the workflow. See Create Sample Report output for details.
- QC & Reports folder:
- Trim Reads report. A report summarizing the performed adapter trimming. See https://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=Trim_Reads.html for details.
- Quantify QIAseq RNA report.
Expression tracks are displayed as tables listing genes included in the panel and several measures of their expression levels.
- Expression value The number of distinct UMIs seen for this gene.
- TPM (Transcripts per Million) The number of transcripts per million that come from this gene. This is computed as the relative abundance per million
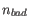 .
.
- RPKM (Reads Per Kilobase per Million reads) There is no good definition of RPKM for targeted amplicon data. We therefore define RPKM to be equal to TPM, which preserves the expected property that RPKM is proportional to TPM.
- Total gene reads The number of distinct UMIs seen for this gene.
- Read counts The total number of reads mapping to this gene. Several reads may have the same UMI.
- UMI counts The number of distinct UMIs seen for this gene.
- Mean reads per UMI The mean number of reads for each UMI seen for this target.
Expression tracks can be further analyzed using statistical tools from the RNA-Seq Analysis folder of the CLC Workbench, such as PCA for RNA-Seq and Create Heat Map for RNA-Seq.
QC metrics can be found in the sample report. They indicate how many reads were ignored and the reason they were not included in a UMI read. The "Accepted reads" column contains the number of reads that passed the filtering carried out by Quantify QIAseq RNA, see Quantify QIAseq RNA.
When you are designing a customized panel online, you will see that there is an option to add a set of 6 "GDC controls". These values are your negative controls of genomic DNA. You can also see these in the target regions where they have names like "GDC_CONTROL_06". The number of control targets with more than 10 UMI reads are listed in the "Expressed GDC controls" table cell. This cell will be colored pink and a warning will be shown if any controls are expressed.
