Output from Detect MSI Status
Three outputs are produced by the Detect MSI Status tool:
- MSI loci track An annotation track with MSI loci annotated with predicted stability.
- MSI report A report summarizing the overall status of the sample and showing length distribution plots of the individual loci.
- Baseline cross-validation report A report analyzing the quality of the MSI baseline.
The MSI report contains both combined and per loci information on stability and other descriptive statistics. The summary section contains information about the number of stable and unstable loci, as well as the MSI status of the sample (figure 8.6).
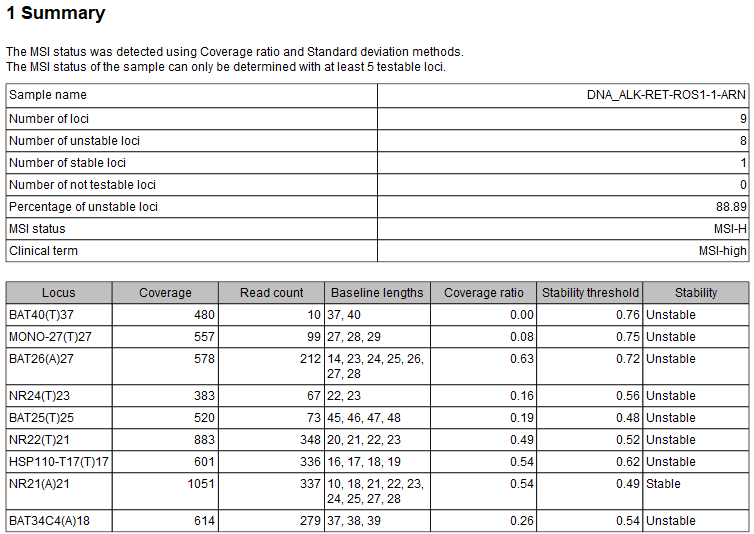
Figure 8.6: Summary section from the MSI Report for an MSI-high sample analyzed using the coverage ratio method.
The loci overview section provides details about the analyzed loci and their stability. The table contains the following information:
- Locus Name of the locus. The name links to a plot showing the distribution of locus lengths for this locus.
- Coverage The number of reads intersecting the locus.
- Read count The number of reads that contains both flanking signatures and have been used for the calculation of the frequency distribution of microsatellite locus lengths as described above.
- Baseline lengths The set of baseline lengths used for determining the stability with coverage ratio and earth mover's distance methods. The column is not shown for the multinomial distribution method.
- Coverage ratio / Earth mover's distance / p-value Stability value from the method of choice, see Detect MSI Status for details on how the metric is calculated.
- Stability threshold / p-value threshold Threshold calculated from the baseline used to assess the stability of the locus. A locus is unstable if the test sample has a stability value:
- below this threshold for the coverage ratio method.
- above this threshold for the earth mover's distance method.
- above this threshold for the multinomial distribution method.
- Stability The stability of a locus can be stable, unstable or N/A. The stability is set to N/A if either the sample or the baseline has insufficient read count.
If the read count is low but the coverage is high, it could be an indication that the locus is highly unstable and only few reads are spanning the locus. Investigating the read mapping can help understanding the problems.
Figure 8.6 shows an example of an MSI report (the loci overview table is truncated by the dashed line), where a sample is compared to the dna_msisensor2_baseline_v1.3 baseline from the Reference Data. The baseline has 120 loci, where two of them are not testable due to too few reads. 107 of the remaining 118 loci are unstable, meaning that the overall assessment of the sample is MSI-high.
The length distribution plot compares the loci lengths observed for the sample (blue) and the baseline (black) for the locus 1_7920926_10[A]. The baseline distribution shows that >90% of the reads have a length of 10 bp, while 85% of reads in the sample have a length of 9 bp. The length distributions are significantly different between the sample and the baseline, and the locus is therefore evaluated as unstable.
The baseline cross-validation report (not shown) contains a table where the MSI status is presented for each sample in the baseline sample set.
The cross-validation analysis verifies whether the baseline and selected parameters are suitable.
For this, the MSI status of each sample from the baseline sample set is tested against a baseline created using all other samples of the set.
Ideally, it is expected that all samples will be detected as stable (MSS) with a very low proportion of unstable loci.
If this is not the case, the parameters might need to be adjusted and/or one or more samples should be removed from the baseline.
Note that the cross-validation analysis is dependent on the parameters used for detection (exactly as for a test sample) and therefore each cross-validation is only valid for the selected set of parameter values used in the cross-validation run.
