Output from the Identify QIAseq DNA Variants workflows
The Identify QIAseq DNA (Pro) Variants workflows produce a Genome Browser View (![]() ) as well as the following files, available in a subfolder (as seen in figure 12.8):
) as well as the following files, available in a subfolder (as seen in figure 12.8):
- A Trim Reads report (
 ) where you can check that adapters were detected by the automatic detection option.
) where you can check that adapters were detected by the automatic detection option.
- A UMI Groups report (
 ) containing a breakdown of UMI groups with different number of reads, along with percentage of groups and reads.
) containing a breakdown of UMI groups with different number of reads, along with percentage of groups and reads.
- A Create UMI report (
 ) that indicates how many reads were ignored and the reason why they were not included in a UMI read.
) that indicates how many reads were ignored and the reason why they were not included in a UMI read.
- A Structural Variants report (
 ) giving an overview of the different types of structural variants inferred by the Structural Variant analysis.
) giving an overview of the different types of structural variants inferred by the Structural Variant analysis.
- A Coverage report (
 ) and a coverage track (
) and a coverage track ( ) from the QC for Target Sequencing tool (see http://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=QC_Targeted_Sequencing.html).
) from the QC for Target Sequencing tool (see http://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=QC_Targeted_Sequencing.html).
- A Sample report (
 ) that contains compiled QC metrics from other reports and provides an overview of a given sample. The sample report also reports whether the QC thresholds specified in the Create Sample Report wizard dialog have been met.
) that contains compiled QC metrics from other reports and provides an overview of a given sample. The sample report also reports whether the QC thresholds specified in the Create Sample Report wizard dialog have been met.
- A read mapping of the UMI Reads (
 ).
).
- Three variant tracks (
 ) : Two from the Variant Caller: the Unfiltered Variants is output before the filtering steps, the Variants passing filters is the one used in the Genome Browser View (for a definition of the variant table content, see http://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=_annotated_variant_table.html). The third is the Indels indirect evidence track produced by the Structural Variant Caller. This is also available in the Genome Browser View.
) : Two from the Variant Caller: the Unfiltered Variants is output before the filtering steps, the Variants passing filters is the one used in the Genome Browser View (for a definition of the variant table content, see http://resources.qiagenbioinformatics.com/manuals/clcgenomicsworkbench/current/index.php?manual=_annotated_variant_table.html). The third is the Indels indirect evidence track produced by the Structural Variant Caller. This is also available in the Genome Browser View.
- An inversion and a long indels track (
 ) containing identified inversions and indels longer than 100,000 bp.
) containing identified inversions and indels longer than 100,000 bp.
- An Amino Acid track (
 )
)
- If a read mapping was submitted in the Copy Number Variant Detection dialog, the workflow will also output three CNV tracks (
 ) (Target-, Region- and Gene-level) and a CNV Results report.
) (Target-, Region- and Gene-level) and a CNV Results report.
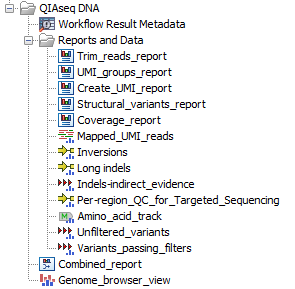
Figure 12.8: Output from the Identify QIAseq DNA (Pro) Variants workflow without CNV detection.
The Unfiltered variant track is included in the output so you can also review why a variant that was expected in the output would have been filtered out of the Variants passing filters track. The difference between the Unfiltered variant track and the Variants passing filters track depends on the following options available in the filtering steps:
- Filter based on quality criteria: Average Quality (quality of the sequenced bases that carry the variant), QUAL (significance of the variant), Read Position Test Probability (relative location of the variant in the reads that cover the variant position - only used in IonTorrent workflows) and Read Direction Test Probability (relative presence of the variant in the reads from different directions that cover the variant position).
- Remove homopolymer error type variants, i.e., errors of the indel type that occur in homopolymer regions. These regions are known to be harder to sequence than non-homopolymeric regions. Note that the definition of homopolymer regions differs between the pipelines due to differences in sequencing technology.
- Remove false positive based on frequency The variant's frequency needs to be above that threshold for the variant to be output by the workflow in the filtered variant track. Note that the unfiltered variant track is generated by the Low Frequency Variant Detection tool run with a frequency cut-off value of 0.5. This value can be considered a pre-filter, which is initially applied to each site in the alignment and determines which sites the variant caller should consider potential variant sites when it starts the error rate and site type/frequencies parameter estimation. In the case of this option, a frequency cut-off is applied on the final candidate variant set (after variants that span across multiple alignment sites have been reconstructed). It is only meaningful to apply this post-filter at a value that is at least as high as the pre-filter value, and we actually recommend using a value that is as least twice as high (1.0). This allows for some wiggle-room when going from the single-site to the multiple site variant construction, in particular to avoid that long InDels are fragmented due to coverage difference throughout the considered region.
The workflow also produces a QC report for the target enrichment that offers statistics on the numbers of targets for which all positions are covered by the "Minimum coverage" threshold set in the QC for targeted sequencing dialog.
The read mapping of the merged UMI groups will let you verify the found variants, and examine why expected variants were not found. The UMI Groups Report gives information about the number of UMI groups found, and how many reads are in each. It includes the following information:
- How many reads were aligned to the reference (Reads in input).
- How many reads were mapped in multiple places and discarded if the Exclude ambiguously mapped reads has been selected, otherwise this value will be shown as zero.
- Groups merged: How many groups were created by merging singleton groups with other groups.
- Number of groups that were discarded for being too small (by default 0 but the option "Minimum group size" of the Calculate Unique Molecular Index Groups can be set up to discard small groups), and how many reads were thus discarded.
- How many groups were created, and of these how many were singletons groups (groups made with sequences sharing identical UMI).
- How many reads are in the largest group.
- How many different UMIs are in the most divergent group (different sequences with different UMIs can be in the same group, if they start on the same position and if they have UMIs that only differ with one character).
- Statistics about the number of reads in the groups.
- Statistics about groups size and reads not included in these groups (also available as graphs below the table).
