Table for Clonotypes
TCR clonotypes (- Clonotype #: A unique number identifying the clonotype.
- Chain:
Which chain type the clonotype belongs to: TRA (
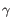 ), TRB (
), TRB (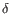 ), TRG (
), TRG (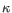 ) and TRD (
) and TRD (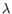 ) for TCR, or IGH (heavy), IGK (light
) for TCR, or IGH (heavy), IGK (light 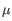 ), and IGL (light
), and IGL (light 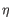 ) for BCR. Note that other light BCR chain types are currently not supported.
) for BCR. Note that other light BCR chain types are currently not supported.
- V, D, J and C: The identified V, D, J and C reference segment(s). If a single unambiguous V / D / J / C segment could not be identified, the segments will be listed separated by a comma.
- CDR3 nucleotide sequence: The nucleotide sequence for CDR3 including the V and J segment-encoded conserved amino acids.
- CDR3 amino acid sequence: The translated amino acid sequence for the CDR3 nucleotide sequence provided that it is in-frame.
- CDR3 length: The length of the CDR3 nucleotide sequence.
- Count: The number of fragments for which the specific clonotype was detected. A fragment represents one single read or one pair of reads.
- Frequency (%): The count given as a percentage relative to the sum of all counts. Note that filtering can affect frequencies, see Filter Immune Repertoire.
- Productive:
One of three categories are used to characterize the CDR3 nucleotide sequences:
- Productive: sequences that are in frame and do not contain a premature stop codon;
- Out-of-frame: sequences that have a length that is not a multiple of three;
- Premature stop codon: sequences that are in-frame but contain a premature stop codon.
The clonotypes are sorted by frequency in decreasing order.
CDR3 quality sore
The CDR3 nucleotide sequences can be colored using the average quality score of each position, by using the "Color CDR3 by average quality sore" option in the Side Panel menu to the right. This option is disabled when there are no quality scores available.
Clonotypes from selection
At the bottom of the table, a button labeled Create Clonotypes from Selection is available. Select the relevant rows in the table and click the button to create new TCR clonotypes (![]() ) or BCR clonotypes (
) or BCR clonotypes (![]() ) that only include the selected clonotypes. When the button is clicked, a dialog with the following options is shown:
) that only include the selected clonotypes. When the button is clicked, a dialog with the following options is shown:
- Recalculate frequencies. If ticked, frequencies in the output clonotypes are recalculated such that they add up to 100% across all chains. Otherwise, the original frequencies found in the input are used.
It can be useful to recalculate frequencies when removing noise (for example, removing clonotypes with a count of 1), but if a subset of clonotypes is created for the purpose of comparing clonotypes between samples, it might be more relevant to preserve the original frequencies.
- Set frequencies per chain. If ticked, the frequencies are recalculated to add up to 100% for each individual chain. This option is enabled only when Recalculate frequencies is ticked.
