Header view
The header view shows details of Genotype track annotations and filters (figure 10.4). The following information is provided:
- Key
- The annotation or filter ID.
- Type
- The annotation value type. Square brackets indicates a list.
- Export tag
- Tag used as ID when the annotation is exported, for example to VCF.
- Category
- Specifies whether it is a sample/database annotation at the locus/variant/alt variant/haplotype allele/haplotype level, or a filter.
- Mode
- Calculation mode of the annotation value:
NOT_CALCULATABLE- Annotation values can only be stored, not calculated.CALCULATE_AND_STORE- Values are calculated and stored when the track is created.CALCULATE_ONTHEFLY_IF_STORED_VALUE_MISSING- If an element has the annotation, calculate the value on-the-fly if there is no stored value, otherwise use the stored value.CALCULATE_ONTHEFLY_NEVER_STORE- If an element has the annotation, calculate the value on-the-fly when requested.
- All annotated
- Are all elements that fit the category considered to have the annotation, or must each annotated element carry the annotation key.
- Importance
- Affects default display of the annotation:
HIGH- annotation will be shown as default in all tables.MEDIUM- annotation will be shown as default in the appropriate table.LOW- annotation will not be shown as default in any table.
- CLC reserved
- Is this a reserved CLC annotation.
- Description
- Description of the annotation or filter.
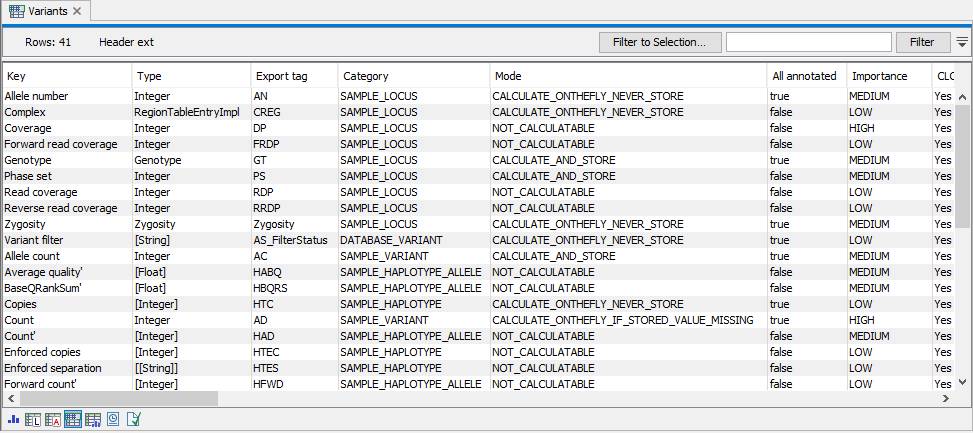
Figure 10.4: The header view. To go to the header view of a genotype track as shown here, select the table icon with a blue table at the top.
