Ploidy and genetic code
Clicking Next offers options for setting ploidy and genetic code (see figure 31.7:
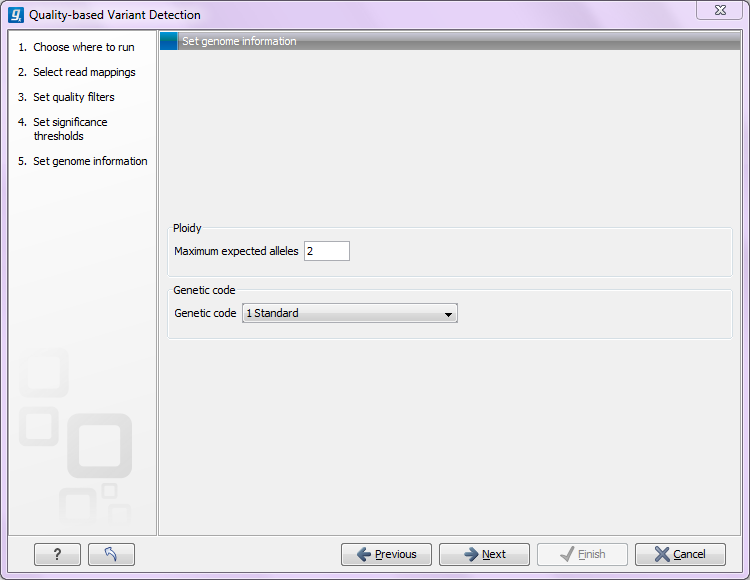
Figure 31.6: Ploidy and genetic code.
- Maximum expected alleles. Allows the user to flag variants that fall in locations with an unexpectedly high number of observed alleles. For a given variant, the entry in the 'hyper-allelic' column of the variant table will contain 'yes', if more than the user-specified 'maximum expected alleles' is observed at the variant position, other observations will result in 'no'.
Note, that with this interpretation the "yes" flag holds true regardless of whether the sequencing data are generated from a population sample or from an individual sample. For example, using a minimum variant frequency of 30% with a diploid organism, you are allowing variants with up to 3 different alleles within the sequencing reads, and by then setting the maximum expected variants count to 2 (the default), any variant with 3 different alleles will be marked as "yes". Genetic code. For the table report, the variant's effect on the protein level is calculated, and the translation table specified here is used. When reporting the variant as a track, this setting has no effect, since the amino acid consequences are calculated separately (see Amino acid changes).
