Reassemble contig
If you have edited a contig, changed trimmed regions, or added or removed reads, you may wish to reassemble the contig. This can be done in two ways:
Toolbox | Molecular Biology Tools (![]() ) | Sequencing Data Analysis (
) | Sequencing Data Analysis (![]() )| Reassemble
Contig (
)| Reassemble
Contig (![]() ) | select the contig from Navigation Area, move to 'Selected Elements' and click
Next
) | select the contig from Navigation Area, move to 'Selected Elements' and click
Next
or right-click in the empty white area of the contig |
Reassemble contig (![]() )
)
This opens a dialog as shown in figure 17.18
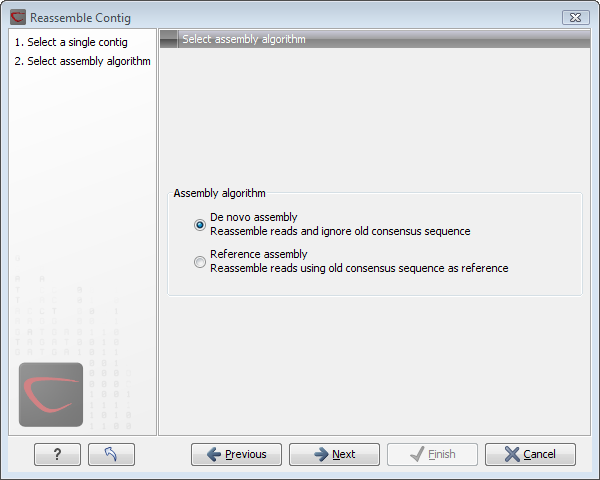
Figure 17.18: Re-assembling a contig.
In this dialog, you can choose:
- De novo assembly. This will perform a normal assembly in the same way as if you had selected the reads as individual sequences. When you click Next, you will follow the same steps as described in Assemble sequences. The consensus sequence of the contig will be ignored.
- Reference assembly. This will use the consensus sequence of the contig as reference. When you click Next, you will follow the same steps as described in Assemble to reference sequence.
When you click Finish, a new contig is created, so you do not lose the information in the old contig.
