Pfam domain search
With CLC Genomics Workbench you can perform a search for domains in protein sequences using the Pfam database. The Pfam database [Bateman et al., 2004] at http://pfam.sanger.ac.uk/ was initially developed to aid the annotation of the C. elegans genome. The database is a large collection of multiple sequence alignments, and contains profile hidden Markov models (HMMs) for individual domain alignments, which can be used to quickly identify domains in protein sequences.
Many proteins have a unique combination of domains, which can be responsible for the catalytic activities of enzymes. Annotating sequences based on pairwise alignment methods by simply transferring annotation from a known protein to the unknown partner does not take domain organization into account [Galperin and Koonin, 1998]. For example, a protein may be annotated incorrectly as an enzyme if the pairwise alignment only finds a regulatory domain.
Using the Pfam Domain Search tool in CLC Genomics Workbench, you can search for domains in sequence data which otherwise do not carry any annotation information. The domain search is performed using the hmmsearch tool from the HMMER3 package version 3.1b1 (http://hmmer.janelia.org/). The Pfam search tool annotates protein sequences with all domains in the Pfam database that have a significant match. It is possible to lower the significance cutoff thresholds in the hmmsearch algorithm, which will reduce the number of domain annotations. Individual domain annotations can be removed manually as described in Removing annotations.
When you have downloaded the Pfam database you are ready to perform a Pfam domain search. To do this start the Pfam search tool:
Toolbox | Classical Sequence Analysis (![]() ) | Protein Analysis (
) | Protein Analysis (![]() )| Pfam Domain Search (
)| Pfam Domain Search (![]() )
)
If a sequence was selected before choosing the Toolbox action, this sequence is now listed in the Selected Elements window of the dialog. Use the arrows to add or remove sequences or sequence lists from the selected elements.
You can perform the analysis on several protein sequences at a time. This will add annotations to all the sequences. Click Next to adjust parameters (see figure 17.10).
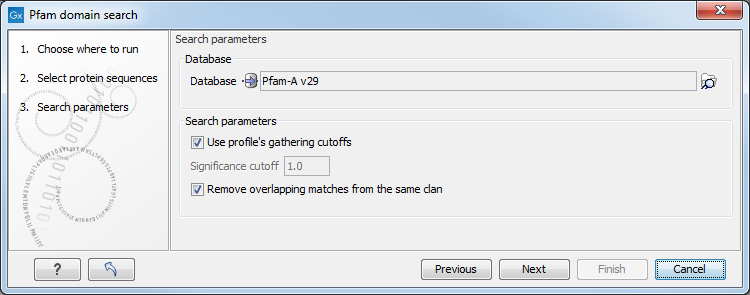
Figure 17.10: Setting parameters for Pfam Domain Search.
- Database. Choose which database to use when searching for Pfam domains. For information on how to download a Pfam database see .
- Significance cutoff
- Use profile's gathering cutoffs. Use cutoffs specifically assigned to each family by the curator instead of manually assigning the Significance cutoff.
- Significance cutoff. The E-value (expectation value) describes the number of hits one would expect to see by chance when searching a database of a particular size. Essentially, a hit with a low E-value is more significant compared to a hit with a high E-value. By lowering the significance threshold the domain search will become more specific and less sensitive, i.e. fewer hits will be reported but the reported hits will be more significant on average.
- Remove overlapping matches from the same clan. Perform post-processing of the results where overlaps between hits are resolved by keeping the hit with the smallest e-value.
Click Next to adjust the output of the tool. The Pfam search tool can produce two types of output. It can add annotations on the input sequences that show the domains found (see figure 17.11) and it can output a table with all the domains found.
Click Finish to start the tool.
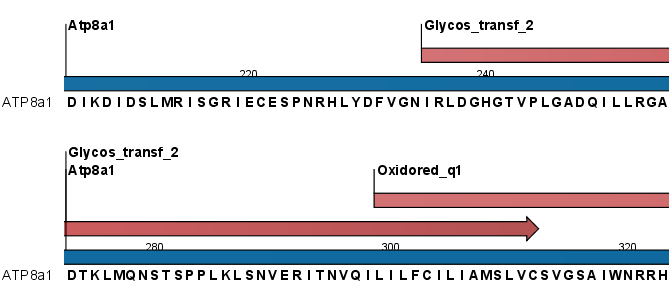
Figure 17.11: Annotations (in red) that were added by the Pfam search tool.
Domain annotations added by the Pfam search tool have the type Region. If the annotations are not visible they have to be enabled in the side panel. Detailed information for each domain annotation, such as the bit score which is the basis for the prediction of domains, is available through the annotation tool tip.
A more detailed description of the scores provided in the annotation tool tips can be found here: http://pfam.sanger.ac.uk/help#tabview=tab5.
