Overlapping paired reads
Depending on the sequencing protocol paired reads may overlap. If the mates of an overlapping pair disagree on one or more positions (e.g. due to sequencing errors) the viewer will represent this as a paired read with either NNN's (gap) or a IUPAC code inserted. This reflects the way the Workbench variant callers would treat disagreements - if two overlapping reads do not agree about the variant base, they are both ignored. If you wish to inspect the mates of overlapping pairs you can check the side panel option 'Disconnect paired reads'. An example is shown in figure 25.26.
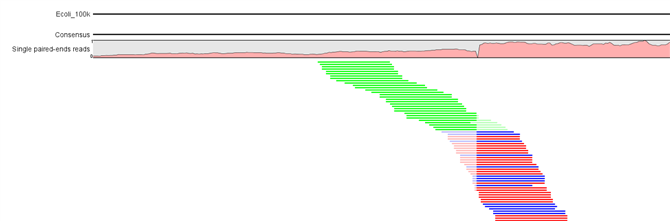
Figure 25.26: Overlapping paired reads containing discrepancies in the overlapping region. The divergent area in the overlapping read pair is marked with N's and IUPAC codes in the top part of the figure. Disconnecting the paired reads by ticking the box "Disconnect Paired Reads" in the Side Panel allows you to see the individual reads.
