Reverse translation from protein into DNA
A protein sequence can be back-translated into DNA using CLC Genomics Workbench. Due to degeneracy of the genetic code every amino acid could translate into several different codons (only 20 amino acids but 64 different codons). Thus, the program offers a number of choices for determining which codons should be used. These choices are explained in this section. For background information see Bioinformatics explained: Reverse translation.
In order to make a reverse translation:
Select a protein sequence | Toolbox in the Menu
Bar | Classical Sequence Analysis (![]() ) | Protein Analysis (
) | Protein Analysis (![]() )|
Reverse Translate (
)|
Reverse Translate (![]() )
)
or right-click a protein sequence | Toolbox |
Classical Sequence Analysis (![]() ) | Protein Analysis (
) | Protein Analysis (![]() )| Reverse
translate (
)| Reverse
translate (![]() )
)
This opens the dialog displayed in figure 16.21:
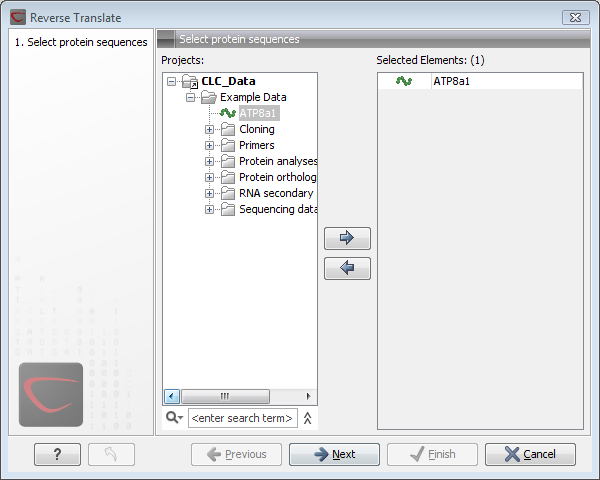
Figure 16.21: Choosing a protein sequence for reverse translation.
If a sequence was selected before choosing the Toolbox action, the sequence is now listed in the Selected Elements window of the dialog. Use the arrows to add or remove sequences or sequence lists from the selected elements. You can translate several protein sequences at a time.
Click Next to adjust the parameters for the translation.
Subsections
