Dynamic motifs
In the Side Panel of sequence views, there is a group called Motifs (see figure 14.24).
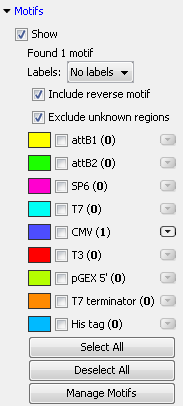
Figure 14.24: Dynamic motifs in the Side Panel.
The Workbench will look for the listed motifs in the sequence that is open and by clicking the check box next to the motif it will be shown in the view as illustrated in figure 14.25.
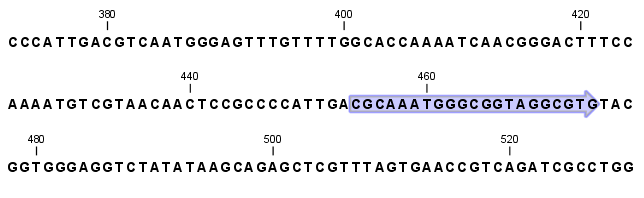
Figure 14.25: Showing dynamic motifs on the sequence.
This case shows the CMV promoter primer sequence which is one of the pre-defined motifs in CLC Genomics Workbench. The motif is per default shown as a faded arrow with no text. The direction of the arrow indicates the strand of the motif.
Placing the mouse cursor on the arrow will display additional information about the motif as illustrated in figure 14.26.

Figure 14.26: Showing dynamic motifs on the sequence.
To add Labels to the motif, select the Flag or Stacked option. They will put the name of the motif as a flag above the sequence. The stacked option will stack the labels when there is more than one motif so that all labels are shown.
Below the labels option there are two options for controlling the way the sequence should be searched for motifs:
- Include reverse motifs. This will also find motifs on the negative strand (only available for nucleotide sequences)
- Exclude matches in N-regions for simple motifs. The motif search handles ambiguous characters in the way that two residues are different if they do not have any residues in common. For example: For nucleotides, N matches any character and R matches A,G. For proteins, X matches any character and Z matches E,Q. Genome sequence often have large regions with unknown sequence. These regions are very often padded with N's. Ticking this checkbox will not display hits found in N-regions and if a one residue in a motif matches to an N, it will be treated as a mismatch.
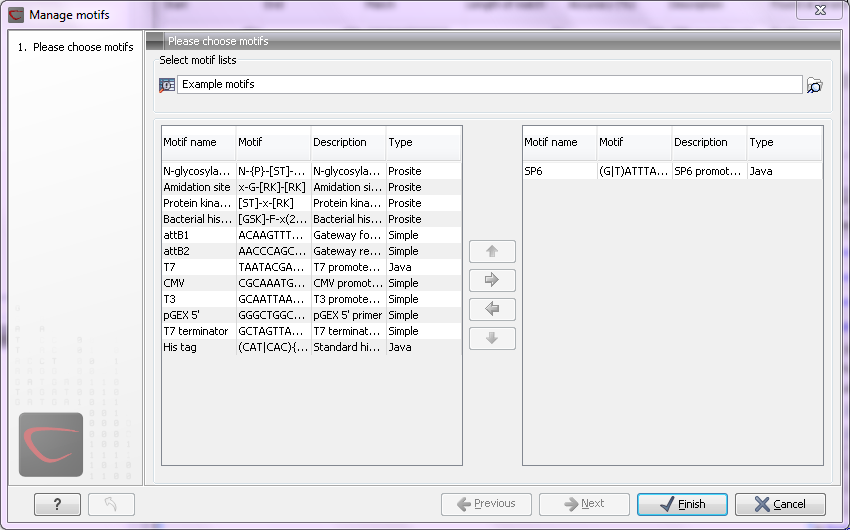
Figure 14.27: Managing the motifs to be shown.
At the top, select a motif list by clicking the Browse (![]() ) button. When the motif list is selected, its motifs are listed in the panel in the left-hand side of the dialog. The right-hand side panel contains the motifs that will be listed in the Side Panel when you click Finish.
) button. When the motif list is selected, its motifs are listed in the panel in the left-hand side of the dialog. The right-hand side panel contains the motifs that will be listed in the Side Panel when you click Finish.
