Identify Shared Variants
This tool should be used if you are interested in finding common (frequent) variants in a group of samples. For example one use case could be that you have 50 unrelated individuals with the same disease and would like to identify variants that are present in at least 70% of all individuals. It can also be used to do an overall comparison between samples (a frequency threshold of 0% will report all alleles).
Toolbox | Resequencing Analysis (![]() ) | Variants Comparison (
) | Variants Comparison (![]() ) | Identify Shared Variants (
) | Identify Shared Variants (![]() )
)
This opens a dialog where you can select the variant tracks (![]() ) from the samples in the group.
) from the samples in the group.
Clicking Next will display the dialog shown in figure 30.9.
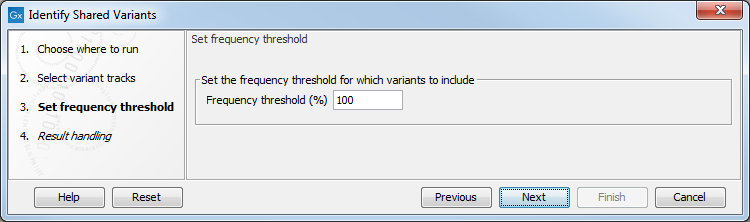
Figure 30.9: Frequency treshold.
The Frequency threshold is the percentage of samples that have this variant. Setting it to 70% means that at least 70% of the samples selected as input have to contain a given variant for it to be reported in the output.
The output of the analysis is a track with all the variants that passed the frequency thresholds and with additional reporting of:
- Sample count. Number of samples that have the variant
- Total number of samples. Total number of samples (this will be identical for all variants).
- Sample frequency. Frequency that is also used as a threshold (see figure 30.9).
- Origin tracks. Comma-separated list of the name of the tracks that contain the variant.
- Homozygous frequency. Percentage of samples passing the filter which have Zygosity annotation homozygous.
- Heterozygous frequency. Percentage of samples passing the filter which have zygosity annotation heterozygous.
- Allele frequency. Mean frequency of the allele in all input samples.
Note that this tool can be used for merging all variants from a number of variant tracks into one track by setting the frequency threshold to 0.
