Element information
The
normal view of a sequence (by double-clicking) shows the annotations
as boxes along the sequence, but often there is more information
available about sequences. This information is available through the
Element info view.
To view the sequence information:
Select a sequence in the Navigation Area and right-click on the file name | Hold the mouse over "Show" to enable a list of options | Element Info (![]() )
)
Another way to show the text view is to open the sequence in the View Area and click on the "Show Element Info" icon (![]() ) found at the bottom of the window.
) found at the bottom of the window.
This will display a view similar to fig 10.16.
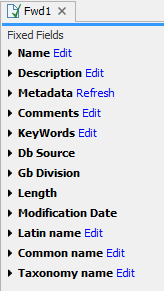
Figure 10.16: The initial display of sequence info for the HUMHBB DNA sequence from the Example data.
All the lines in the view are headings, and the corresponding text can be shown by clicking the text. The information available depends on the origin of the sequence.
- Name. The name of the sequence which is also shown in sequence views and in the Navigation Area.
- Description. A description of the sequence.
- Metadata. The Metadata table and the detailed metadata values associated with the sequence.
- Comments. The author's comments about the sequence.
- Keywords. Keywords describing the sequence.
- Db source. Accession numbers in other databases concerning the same sequence.
- Gb Division. Abbreviation of GenBank divisions. See section 3.3 in the GenBank release notes for a full list of GenBank divisions.
- Length. The length of the sequence.
- Modification date. Modification date from the database. This means that this date does not reflect your own changes to the sequence. See the section on history views for information about the latest changes to the sequence after it was downloaded from the database.
- Latin name. Latin name of the organism.
- Common name. Scientific name of the organism.
- Taxonomy name. Taxonomic classification levels.
- Read group Read group identifier "ID", technology used to produced the reads "Platform", and sample name "Sample".
- Paired Status. Unpaired or Paired sequences, with in this case the Minimum and Maximum distances as well as the Read orientation set during import.
Some of the information can be edited by clicking the blue Edit text. This means that you can add your own information to sequences that do not derive from databases.
